Genomics
This section list compuational tools and web server in the field of genomics. This includes genome annotation, genome-based medicine, Cancer genomics and genome based biomarkers.
This section host servers that is important for annotating genome (or nucleotide) sequences (directly or indirectly). These tools cover wide range of applications that includes; i) genome wide similarity search, ii) repeats in nucleotide sequence, iii) identification of protein coding regions or genes, iv) classification of RNA families and v) designing siRNA and miRNA.
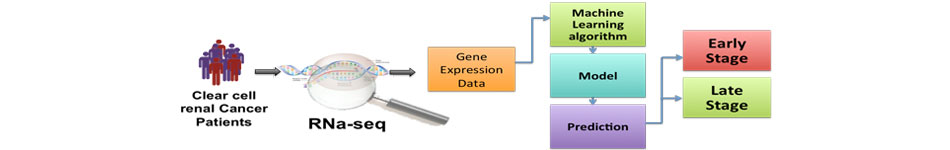
One of major challenges in handling cancer is to annotate big data in the the field of cancer genomics. Aim of this module is to provide in silico solutions for cancer biologists. This page manitain tools and databases developed in the field of cancer informatics, it includes; i) drug resistance databases, ii) genome based cancer biomarkers (e.g., stage classification), iii) epitope-based vaccines, iv) databases.
Personalised medicine is a new concept in the field of drug or vaccine design. In this module our focus is to utilise genomic or genetic information for designing medicine against disease like cancer, tuberculosis. This page maintain wide range of computational resources, like i) prioritization of drugs based on genomic information, ii) personalised vaccine agains cancer, iii) strain-specific vaccines against bacteria.
One of the major challenges in the field oh health sciences is to design genome based biomarkers, as these biomarkers are more sensitive and accurate than traditional tests. Here, aim is to develop computational tools to predict wide range of biomrakers from genomic information. These resources can be classified in following categories; i) disease identification, ii) disease progression, iii) drug biomarkers.