Antiviral vaccines have been the most successful biomedical intervention for preventing epidemic viral disease. Vaccination for smallpox in humans was the first successful example for the eradication of the disease. While early vaccines were developed empirically by passage in live animals or eggs, more recent vaccines have been developed because of the advent of new technologies, particularly cell culture and molecular biology. Technological advances in gene delivery and expression, nanoparticles, protein manufacturing, and adjuvants have created the potential for new vaccine platforms that may provide solutions for vaccines against viral pathogens for which no interventions currently exist. In recent years, use of In silico methods for designing and prioritizing vaccine candidates becomes the backbone of the pharmaceutical industries. This draws the attention of the researchers and scientists working in this field.
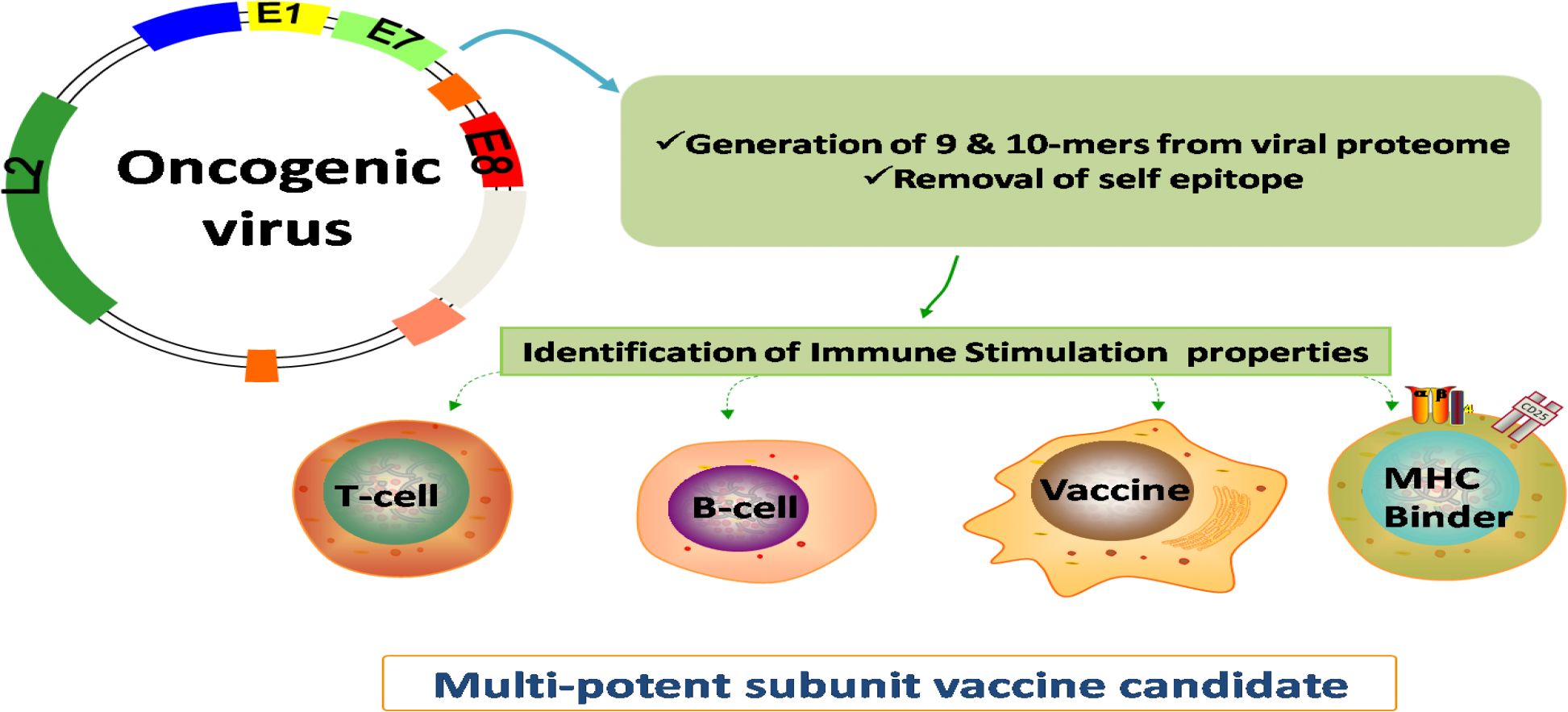
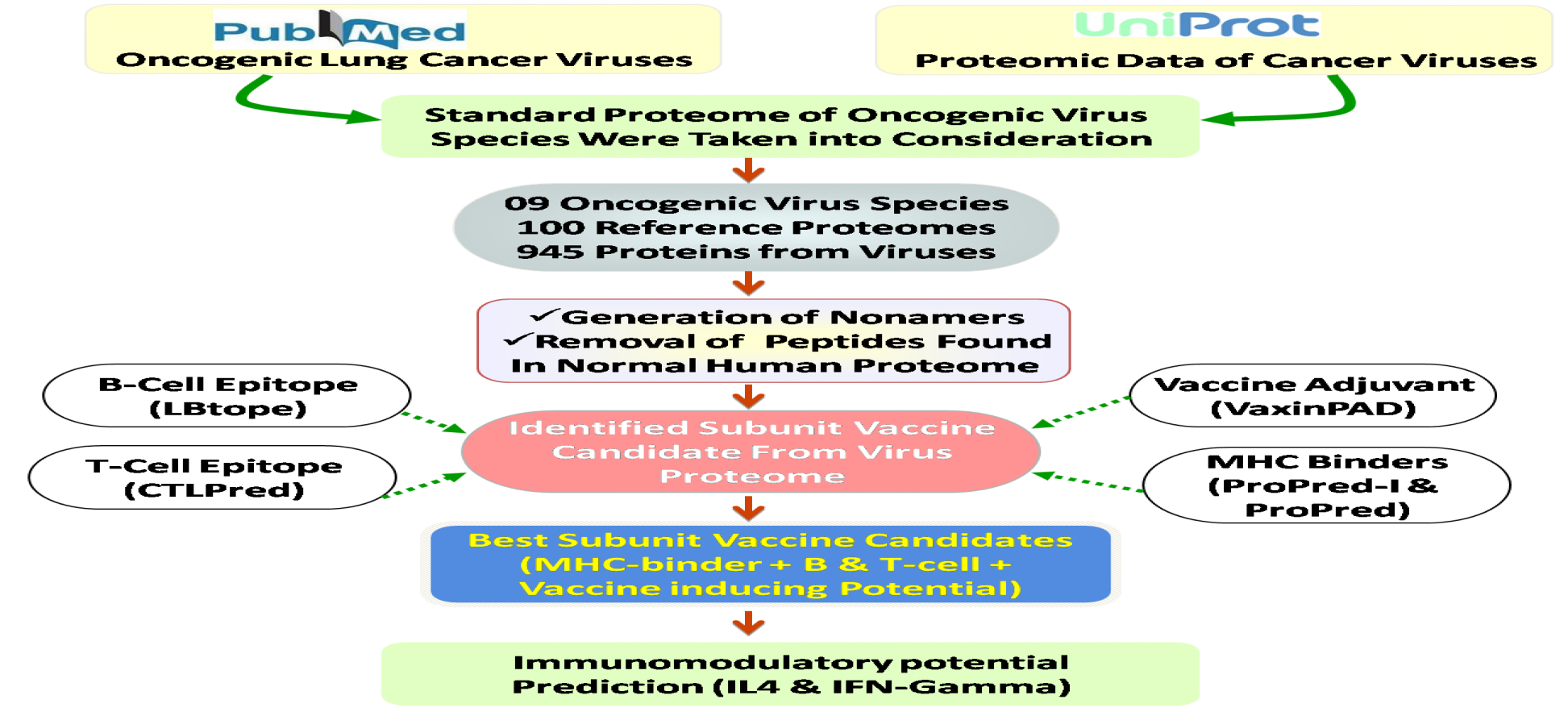
The present platform is designed to guide the vaccine designing process against major lung cancer causing oncogenic viruses. The genomic and proteomic information of 09 oncogenic lung cancer virus species, 100 reference proteome of virus strains as well as 945 target proteins information was taken into consideration. The identified vaccine candidates have the potential to stimulate adaptive immunity, thus helps in providing both protective and antiviral immunity. We hope the developed platform could best serves the need of scientific community for improving the cancer immunotherapy process.