Methylation Techinque Browse Result Page |
| Search: | ||||||||||||||||||||||||
| PCMDB ID | Origin | Name | Expression of important Genes |
Gene | Cancer Type | Control | Methylation Category |
Mehtylation Value |
Method | Reference | ||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| PCM055435 | Tissue Sample | Sample 29 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055436 | Tissue Sample | Sample 14 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055437 | Tissue Sample | Sample 15 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055438 | Tissue Sample | Sample 18 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055439 | Tissue Sample | Sample 20 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055440 | Tissue Sample | Sample 25 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055441 | Tissue Sample | Sample 26 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055442 | Tissue Sample | Sample 27 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055443 | Tissue Sample | Sample 16 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055444 | Tissue Sample | Sample 21 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055445 | Tissue Sample | Sample 28 | NA | THBS1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055446 | Tissue Sample | Sample 29 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055447 | Tissue Sample | Sample 14 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055448 | Tissue Sample | Sample 15 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055449 | Tissue Sample | Sample 18 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055450 | Tissue Sample | Sample 20 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055451 | Tissue Sample | Sample 25 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055452 | Tissue Sample | Sample 26 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055453 | Tissue Sample | Sample 27 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055454 | Tissue Sample | Sample 16 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055455 | Tissue Sample | Sample 21 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055456 | Tissue Sample | Sample 28 | NA | MGMT | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055457 | Tissue Sample | Sample 29 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055458 | Tissue Sample | Sample 14 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055459 | Tissue Sample | Sample 15 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055460 | Tissue Sample | Sample 18 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055461 | Tissue Sample | Sample 20 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055462 | Tissue Sample | Sample 25 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055463 | Tissue Sample | Sample 26 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055464 | Tissue Sample | Sample 27 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055465 | Tissue Sample | Sample 16 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055466 | Tissue Sample | Sample 21 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055467 | Tissue Sample | Sample 28 | NA | MINT31 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055468 | Tissue Sample | Sample 29 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055469 | Tissue Sample | Sample 14 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055470 | Tissue Sample | Sample 15 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055471 | Tissue Sample | Sample 18 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055472 | Tissue Sample | Sample 20 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055473 | Tissue Sample | Sample 25 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055474 | Tissue Sample | Sample 26 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055475 | Tissue Sample | Sample 27 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055476 | Tissue Sample | Sample 16 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055477 | Tissue Sample | Sample 21 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055478 | Tissue Sample | Sample 28 | NA | MINT27 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055479 | Tissue Sample | Sample 29 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055480 | Tissue Sample | Sample 14 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055481 | Tissue Sample | Sample 15 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055482 | Tissue Sample | Sample 18 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055483 | Tissue Sample | Sample 20 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055484 | Tissue Sample | Sample 25 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055485 | Tissue Sample | Sample 26 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055486 | Tissue Sample | Sample 27 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055487 | Tissue Sample | Sample 16 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055488 | Tissue Sample | Sample 21 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055489 | Tissue Sample | Sample 28 | NA | MINT1 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055490 | Tissue Sample | Sample 29 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055491 | Tissue Sample | Sample 14 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055492 | Tissue Sample | Sample 15 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055493 | Tissue Sample | Sample 18 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055494 | Tissue Sample | Sample 20 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055495 | Tissue Sample | Sample 25 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055496 | Tissue Sample | Sample 26 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055497 | Tissue Sample | Sample 27 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055498 | Tissue Sample | Sample 16 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055499 | Tissue Sample | Sample 21 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055500 | Tissue Sample | Sample 28 | NA | COX2 | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055501 | Tissue Sample | Sample 29 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055502 | Tissue Sample | Sample 14 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055503 | Tissue Sample | Sample 15 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055504 | Tissue Sample | Sample 18 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055505 | Tissue Sample | Sample 20 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055506 | Tissue Sample | Sample 25 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055507 | Tissue Sample | Sample 26 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055508 | Tissue Sample | Sample 27 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055509 | Tissue Sample | Sample 16 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055510 | Tissue Sample | Sample 21 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055511 | Tissue Sample | Sample 28 | NA | RARA | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055512 | Tissue Sample | Sample 29 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055513 | Tissue Sample | Sample 14 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055514 | Tissue Sample | Sample 15 | NA | ER | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055515 | Tissue Sample | Sample 18 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055516 | Tissue Sample | Sample 20 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055517 | Tissue Sample | Sample 25 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055518 | Tissue Sample | Sample 26 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055519 | Tissue Sample | Sample 27 | NA | ER | Endo | normal pancreas adjacent to tumor | Not Known | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055520 | Tissue Sample | Sample 16 | NA | ER | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055521 | Tissue Sample | Sample 21 | NA | ER | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM055522 | Tissue Sample | Sample 28 | NA | ER | Endo | normal pancreas adjacent to tumor | Low | NA | Bisulphite Sequencin | 12584572 | ||||||||||||||
| PCM056974 | Tissue Sample | Sample 1 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056975 | Tissue Sample | Sample 2 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056976 | Tissue Sample | Sample 3 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Low | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056977 | Tissue Sample | Sample 4 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056978 | Tissue Sample | Sample 5 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | High | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056979 | Tissue Sample | Sample 6 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056980 | Tissue Sample | Sample 7 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Low | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056981 | Tissue Sample | Sample 8 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056982 | Tissue Sample | Sample 9 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056983 | Tissue Sample | Sample 10 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056984 | Tissue Sample | Sample 11 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056985 | Tissue Sample | Sample 12 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Low | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056986 | Tissue Sample | Sample 13 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056987 | Tissue Sample | Sample 14 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | High | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056988 | Tissue Sample | Sample 15 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056989 | Tissue Sample | Sample 16 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056990 | Tissue Sample | Sample 17 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056991 | Tissue Sample | Sample 18 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056992 | Tissue Sample | Sample 19 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056993 | Tissue Sample | Sample 20 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056994 | Tissue Sample | Sample 21 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056995 | Tissue Sample | Sample 22 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056996 | Tissue Sample | Sample 23 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056997 | Tissue Sample | Sample 24 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM056998 | Tissue Sample | Sample 25 | NA | p16 | S_not | 25 paired normal tissue away from the main tumor mass | Intermediate | NA | Bisulphite Sequencin | 15985168 | ||||||||||||||
| PCM057899 | Tissue Sample | Tissue no. 117 | NA | p15 | Endo | Normal pancreatic tissue | Not Known | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057900 | Tissue Sample | Tissue no. 5735 | NA | p15 | Endo | Normal pancreatic tissue | Not Known | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057901 | Tissue Sample | Tissue no. 3945 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057902 | Tissue Sample | Tissue No. 4642 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057903 | Tissue Sample | Tissue No. 4741 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057904 | Tissue Sample | Tissue No. 5288 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057905 | Tissue Sample | Tissue No. 6114 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057906 | Tissue Sample | Tissue No. 2643 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057907 | Tissue Sample | Tissue No. 2663 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057908 | Tissue Sample | Tissue No. 5620 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057909 | Tissue Sample | Tissue No. 5625 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057910 | Tissue Sample | Tissue No. 2576 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057911 | Tissue Sample | Tissue No. 3219 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057912 | Tissue Sample | Tissue No. 5516 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057913 | Tissue Sample | Tissue No. 5465 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM057914 | Tissue Sample | Tissue No. 6370 | NA | p15 | Endo | Normal pancreatic tissue | Low | NA | Bisulphite Sequencin | 17803708 | ||||||||||||||
| PCM058275 | Tissue Sample | Patient 1 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058276 | Tissue Sample | Patient 2 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058277 | Tissue Sample | Patient 3 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058278 | Tissue Sample | Patient 4 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058279 | Tissue Sample | Patient 5 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058280 | Tissue Sample | Patient 6 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058281 | Tissue Sample | Patient 7 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058282 | Tissue Sample | Patient 8 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058283 | Tissue Sample | Patient 9 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058284 | Tissue Sample | Patient 10 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058285 | Tissue Sample | Patient 11 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058286 | Tissue Sample | Patient 12 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058287 | Tissue Sample | Patient 13 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058288 | Tissue Sample | Patient 14 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058289 | Tissue Sample | Patient 15 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058290 | Tissue Sample | Patient 16 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058291 | Tissue Sample | Patient 17 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058292 | Tissue Sample | Patient 18 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058293 | Tissue Sample | Patient 19 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058294 | Tissue Sample | Patient 20 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058295 | Tissue Sample | Patient 21 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058296 | Tissue Sample | Patient 22 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058297 | Tissue Sample | Patient 23 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058298 | Tissue Sample | Patient 24 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058299 | Tissue Sample | Patient 25 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058300 | Tissue Sample | Patient 26 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058301 | Tissue Sample | Patient 27 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058302 | Tissue Sample | Patient 28 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058303 | Tissue Sample | Patient 29 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058304 | Tissue Sample | Patient 30 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Low | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058305 | Tissue Sample | Patient 31 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM058306 | Tissue Sample | Patient 32 | NA | RUNX3 | S_not | 32 corresponding non-cancerous tissue | Not Known | NA | Bisulphite Sequencin | 18475302 | ||||||||||||||
| PCM08 | Cell Line | HPAFII | 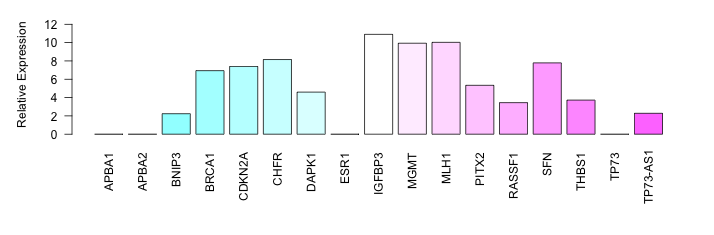  | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM09 | Cell Line | CAPAN2 | 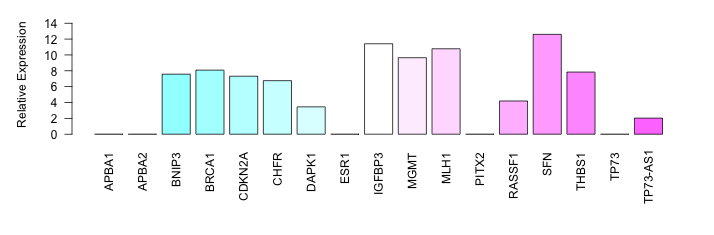  | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM010 | Cell Line | ASPC1 | 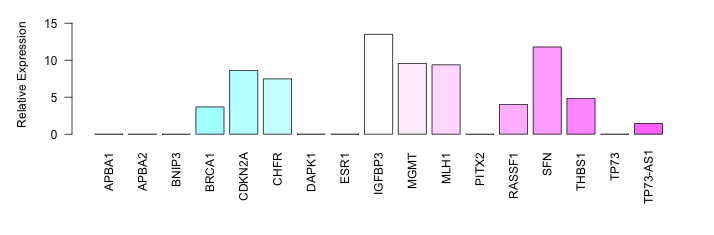  | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM011 | Cell Line | BXPC3 |   | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM012 | Cell Line | MIAPACA2 |   | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM013 | Cell Line | PANC1 |   | NDRG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM014 | Cell Line | BXPC3 |   | TNFRSF10C | S_not | NA | High | 68.84±8.71% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM015 | Cell Line | PANC1 |   | TNFRSF10C | S_not | NA | High | 96.77±4.57% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM016 | Cell Line | SW1990 | 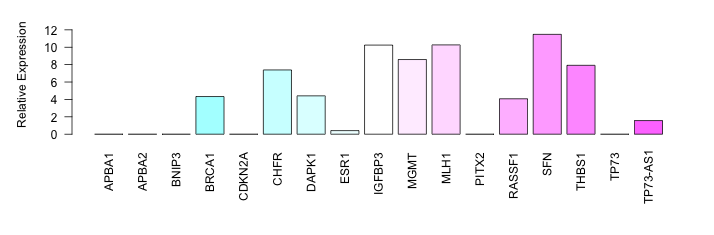  | TNFRSF10C | S_not | NA | High | 54.97±7.33% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM017 | Cell Line | CFPAC1 |   | TNFRSF10C | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM018 | Cell Line | PANC1 |   | TNFRSF10C | S_not | NA | High | 98.00% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM019 | Cell Line | CFPAC1 |   | TNFRSF10C | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM060 | Cell Line | HPAFII |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM061 | Cell Line | HPAFII |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM062 | Cell Line | CAPAN2 |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM063 | Cell Line | ASPC1 |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM064 | Cell Line | BXPC3 |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM065 | Cell Line | MIAPACA2 |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM066 | Cell Line | PANC1 |   | NRDG1 | S_not | NA | Low | NA | Bisulphite Sequencin | 20173668 | ||||||||||||||
| PCM0202 | Cell Line | A818 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0203 | Cell Line | ASPC1 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0204 | Cell Line | BXPC3 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0205 | Cell Line | CAPAN1 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0206 | Cell Line | CAPAN2 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0207 | Cell Line | HPAF2 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0208 | Cell Line | HS766T |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0209 | Cell Line | MPANC96 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0210 | Cell Line | PATU89885 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0211 | Cell Line | PL45 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0212 | Cell Line | SU8686 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0213 | Cell Line | SUIT0028 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0214 | Cell Line | SUIT007 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0215 | Cell Line | PANC1 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0216 | Cell Line | MIAPACA2 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0217 | Cell Line | PATU8902 |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0218 | Cell Line | PATU8988T |   | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0219 | Cell Line | PT45 | NA | AKAP12 | Adeno | NA | Not Known | 83.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0220 | Cell Line | A818 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0221 | Cell Line | ASPC1 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0222 | Cell Line | BXPC3 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0223 | Cell Line | CAPAN1 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0224 | Cell Line | CAPAN2 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0225 | Cell Line | HPAF2 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0226 | Cell Line | HS766T |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0227 | Cell Line | MPANC96 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0228 | Cell Line | PATU89885 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0229 | Cell Line | PL45 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0230 | Cell Line | SU8686 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0231 | Cell Line | SUIT0028 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0232 | Cell Line | SUIT007 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0233 | Cell Line | PANC1 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0234 | Cell Line | MIAPACA2 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0235 | Cell Line | PATU8902 |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0236 | Cell Line | PATU8988T |   | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0237 | Cell Line | PT45 | NA | SERPINB5 | Adeno | NA | Not Known | 28.00% | Bisulphite Sequencin | 20939879 | ||||||||||||||
| PCM0238 | Cell Line | PANC1 |   | ELAB3 | S_not | NA | High | NA | Bisulphite Sequencin | 20428826 | ||||||||||||||
| PCM0239 | Cell Line | PATU8988 | NA | ELAB3 | S_not | NA | High | NA | Bisulphite Sequencin | 20428826 | ||||||||||||||
| PCM0240 | Cell Line | BXPC3 |   | ELAB3 | S_not | NA | High | NA | Bisulphite Sequencin | 20428826 | ||||||||||||||
| PCM0241 | Cell Line | SWI990 | NA | ELAB3 | S_not | NA | High | NA | Bisulphite Sequencin | 20428826 | ||||||||||||||
| PCM0242 | Cell Line | CFPAC | NA | ELAB3 | S_not | NA | High | NA | Bisulphite Sequencin | 20428826 | ||||||||||||||
| PCM0243 | Cell Line | ASPC | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0244 | Cell Line | CAP2 | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0245 | Cell Line | HPAC |   | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0246 | Cell Line | HPAF | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0247 | Cell Line | HS766T |   | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0248 | Cell Line | MIAPACA | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0249 | Cell Line | PANC | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0250 | Cell Line | RWP1 | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0251 | Cell Line | SW1990 |   | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0252 | Cell Line | UCVA | NA | OSMR | S_not | NA | Low | 10.00% | Bisulphite Sequencin | 19223499 | ||||||||||||||
| PCM0256 | Cell Line | BXP3 | NA | STC2 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 18394600 | ||||||||||||||
| PCM0283 | Cell Line | MIAPACA2 |   | Sod2 | S_not | NA | High | NA | Bisulphite Sequencin | 17895890 | ||||||||||||||
| PCM0284 | Cell Line | ASPC1 |   | Sod2 | S_not | NA | Intermediate | NA | Bisulphite Sequencin | 17895890 | ||||||||||||||
| PCM0285 | Cell Line | BXPC3 |   | Sod2 | S_not | NA | Intermediate | NA | Bisulphite Sequencin | 17895890 | ||||||||||||||
| PCM0286 | Cell Line | CAPAN1 |   | Sod2 | S_not | NA | Low | NA | Bisulphite Sequencin | 17895890 | ||||||||||||||
| PCM0357 | Cell Line | MIAPACA2 |   | TCF2 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 16479257 | ||||||||||||||
| PCM0358 | Cell Line | KP3 |   | TCF2 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 16479257 | ||||||||||||||
| PCM0359 | Cell Line | KP1NC | NA | TCF2 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 16479257 | ||||||||||||||
| PCM0360 | Cell Line | CFPAC | NA | TCF2 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 16479257 | ||||||||||||||
| PCM0365 | Cell Line | MIAPACA2 |   | DUSP6 | S_not | NA | High | NA | Bisulphite Sequencin | 15824892 | ||||||||||||||
| PCM0366 | Cell Line | PAN07JCK | NA | DUSP6 | S_not | NA | Intermediate | NA | Bisulphite Sequencin | 15824892 | ||||||||||||||
| PCM0367 | Cell Line | ASPC1 |   | DUSP6 | S_not | NA | Low | NA | Bisulphite Sequencin | 15824892 | ||||||||||||||
| PCM0368 | Cell Line | PANC1 |   | MUC2 CpG1 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0369 | Cell Line | PANC1 |   | MUC2 CpG2 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0370 | Cell Line | PANC1 |   | MUC2 CpG3 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0371 | Cell Line | PANC1 |   | MUC2 CpG4 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0372 | Cell Line | PANC1 |   | MUC2 CpG5 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0373 | Cell Line | PANC1 |   | MUC2 CpG6 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0374 | Cell Line | PANC1 |   | MUC2 CpG7 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0375 | Cell Line | PANC1 |   | MUC2 CpG8 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0376 | Cell Line | PANC1 |   | MUC2 CpG9 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0377 | Cell Line | PANC1 |   | MUC2 CpG10 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0378 | Cell Line | PANC1 |   | MUC2 CpG11 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0379 | Cell Line | PANC1 |   | MUC2 CpG12 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0380 | Cell Line | PANC1 |   | MUC2 CpG13 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0381 | Cell Line | PANC1 |   | MUC2 CpG14 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0382 | Cell Line | PANC1 |   | MUC2 CpG15 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0383 | Cell Line | PANC1 |   | MUC2 CpG16 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0384 | Cell Line | PANC1 |   | MUC2 CpG17 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0385 | Cell Line | PANC1 |   | MUC2 CpG18 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0386 | Cell Line | PANC1 |   | MUC2 CpG19 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0387 | Cell Line | PANC1 |   | MUC2 CpG20 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0388 | Cell Line | PANC1 |   | MUC2 CpG21 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0389 | Cell Line | PANC1 |   | MUC2 CpG22 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0390 | Cell Line | PANC1 |   | MUC2 CpG23 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0391 | Cell Line | PANC1 |   | MUC2 CpG24 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0392 | Cell Line | PANC1 |   | MUC2 CpG25 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0393 | Cell Line | PANC1 |   | MUC2 CpG26 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0394 | Cell Line | PANC1 |   | MUC2 CpG27 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0395 | Cell Line | PANC1 |   | MUC2 CpG28 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0396 | Cell Line | PANC1 |   | MUC2 CpG29 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0397 | Cell Line | PANC1 |   | MUC2 CpG30 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0398 | Cell Line | PANC1 |   | MUC2 CpG31 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0399 | Cell Line | PANC1 |   | MUC2 CpG32 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0400 | Cell Line | PANC1 |   | MUC2 CpG33 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0401 | Cell Line | PANC1 |   | MUC2 CpG34 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0402 | Cell Line | PANC1 |   | MUC2 CpG35 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0403 | Cell Line | PANC1 |   | MUC2 CpG36 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0404 | Cell Line | PANC1 |   | MUC2 CpG37 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0405 | Cell Line | PANC1 |   | MUC2 CpG38 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0406 | Cell Line | PANC1 |   | MUC2 CpG39 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0407 | Cell Line | PANC1 |   | MUC2 CpG40 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0408 | Cell Line | PANC1 |   | MUC2 CpG41 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0409 | Cell Line | PANC1 |   | MUC2 CpG42 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0410 | Cell Line | PANC1 |   | MUC2 CpG43 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0411 | Cell Line | PANC1 |   | MUC2 CpG44 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0412 | Cell Line | PANC1 |   | MUC2 CpG45 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0413 | Cell Line | PANC1 |   | MUC2 CpG46 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0414 | Cell Line | PANC1 |   | MUC2 CpG47 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0415 | Cell Line | PANC1 |   | MUC2 CpG48 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0416 | Cell Line | PANC1 |   | MUC2 CpG49 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0417 | Cell Line | PANC1 |   | MUC2 CpG50 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0418 | Cell Line | PANC1 |   | MUC2 CpG51 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0419 | Cell Line | PANC1 |   | MUC2 CpG52 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0420 | Cell Line | PANC1 |   | MUC2 CpG53 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0421 | Cell Line | PANC1 |   | MUC2 CpG54 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0422 | Cell Line | PANC1 |   | MUC2 CpG55 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0423 | Cell Line | PANC1 |   | MUC2 CpG56 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0424 | Cell Line | PANC1 |   | MUC2 CpG57 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0425 | Cell Line | PANC1 |   | MUC2 CpG58 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0426 | Cell Line | PANC1 |   | MUC2 CpG59 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0427 | Cell Line | BXPC3 |   | MUC2 CpG1 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0428 | Cell Line | BXPC3 |   | MUC2 CpG2 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0429 | Cell Line | BXPC3 |   | MUC2 CpG3 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0430 | Cell Line | BXPC3 |   | MUC2 CpG4 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0431 | Cell Line | BXPC3 |   | MUC2 CpG5 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0432 | Cell Line | BXPC3 |   | MUC2 CpG6 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0433 | Cell Line | BXPC3 |   | MUC2 CpG7 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0434 | Cell Line | BXPC3 |   | MUC2 CpG8 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0435 | Cell Line | BXPC3 |   | MUC2 CpG9 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0436 | Cell Line | BXPC3 |   | MUC2 CpG10 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0437 | Cell Line | BXPC3 |   | MUC2 CpG11 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0438 | Cell Line | BXPC3 |   | MUC2 CpG12 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0439 | Cell Line | BXPC3 |   | MUC2 CpG13 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0440 | Cell Line | BXPC3 |   | MUC2 CpG14 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0441 | Cell Line | BXPC3 |   | MUC2 CpG15 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0442 | Cell Line | BXPC3 |   | MUC2 CpG16 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0443 | Cell Line | BXPC3 |   | MUC2 CpG17 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0444 | Cell Line | BXPC3 |   | MUC2 CpG18 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0445 | Cell Line | BXPC3 |   | MUC2 CpG19 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0446 | Cell Line | BXPC3 |   | MUC2 CpG20 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0447 | Cell Line | BXPC3 |   | MUC2 CpG21 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0448 | Cell Line | BXPC3 |   | MUC2 CpG22 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0449 | Cell Line | BXPC3 |   | MUC2 CpG23 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0450 | Cell Line | BXPC3 |   | MUC2 CpG24 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0451 | Cell Line | BXPC3 |   | MUC2 CpG25 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0452 | Cell Line | BXPC3 |   | MUC2 CpG26 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0453 | Cell Line | BXPC3 |   | MUC2 CpG27 | S_not | Normal colonic crypts | High | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0454 | Cell Line | BXPC3 |   | MUC2 CpG28 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0455 | Cell Line | BXPC3 |   | MUC2 CpG29 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0456 | Cell Line | BXPC3 |   | MUC2 CpG30 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0457 | Cell Line | BXPC3 |   | MUC2 CpG31 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0458 | Cell Line | BXPC3 |   | MUC2 CpG32 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0459 | Cell Line | BXPC3 |   | MUC2 CpG33 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0460 | Cell Line | BXPC3 |   | MUC2 CpG34 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0461 | Cell Line | BXPC3 |   | MUC2 CpG35 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0462 | Cell Line | BXPC3 |   | MUC2 CpG36 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0463 | Cell Line | BXPC3 |   | MUC2 CpG37 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0464 | Cell Line | BXPC3 |   | MUC2 CpG38 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0465 | Cell Line | BXPC3 |   | MUC2 CpG39 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0466 | Cell Line | BXPC3 |   | MUC2 CpG40 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0467 | Cell Line | BXPC3 |   | MUC2 CpG41 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0468 | Cell Line | BXPC3 |   | MUC2 CpG42 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0469 | Cell Line | BXPC3 |   | MUC2 CpG43 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0470 | Cell Line | BXPC3 |   | MUC2 CpG44 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0471 | Cell Line | BXPC3 |   | MUC2 CpG45 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0472 | Cell Line | BXPC3 |   | MUC2 CpG46 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0473 | Cell Line | BXPC3 |   | MUC2 CpG47 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0474 | Cell Line | BXPC3 |   | MUC2 CpG48 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0475 | Cell Line | BXPC3 |   | MUC2 CpG49 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0476 | Cell Line | BXPC3 |   | MUC2 CpG50 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0477 | Cell Line | BXPC3 |   | MUC2 CpG51 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0478 | Cell Line | BXPC3 |   | MUC2 CpG52 | S_not | Normal colonic crypts | Intermediate | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0479 | Cell Line | BXPC3 |   | MUC2 CpG53 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0480 | Cell Line | BXPC3 |   | MUC2 CpG54 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0481 | Cell Line | BXPC3 |   | MUC2 CpG55 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0482 | Cell Line | BXPC3 |   | MUC2 CpG56 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0483 | Cell Line | BXPC3 |   | MUC2 CpG57 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0484 | Cell Line | BXPC3 |   | MUC2 CpG58 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0485 | Cell Line | BXPC3 |   | MUC2 CpG59 | S_not | Normal colonic crypts | Low | NA | Bisulphite Sequencin | 16112420 | ||||||||||||||
| PCM0486 | Cell Line | ASPC1 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0487 | Cell Line | BXPC3 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0488 | Cell Line | CAPAN1 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0489 | Cell Line | CAPAN2 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0490 | Cell Line | CFPAC1 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0491 | Cell Line | MIAPACA2 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0492 | Cell Line | PANC1 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0493 | Cell Line | SU8686 |   | CXCR4 | S_not | NA | Altered | 45.00% | Bisulphite Sequencin | 15662133 | ||||||||||||||
| PCM0494 | Cell Line | PANC1 |   | BNIP3 | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0495 | Cell Line | BXPC3 |   | BNIP3 | S_not | NA | High | 100.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0496 | Cell Line | MIAPACA2 |   | BNIP3 | S_not | NA | High | 100.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0497 | Cell Line | KP3 |   | BNIP3 | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0498 | Cell Line | KPINL | NA | BNIP3 | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0499 | Cell Line | CFPAC | NA | BNIP3 | S_not | NA | Low | 0.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0500 | Cell Line | PK45H |   | BNIP3 | S_not | NA | Intermediate | 68.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0501 | Cell Line | PK59 |   | BNIP3 | S_not | NA | High | 100.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0502 | Cell Line | KLM1 |   | BNIP3 | S_not | NA | High | 91.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0503 | Cell Line | CAPAN2 |   | BNIP3 | S_not | NA | Intermediate | 54.00% | Bisulphite Sequencin | 15942716 | ||||||||||||||
| PCM0504 | Cell Line | ASPC1 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0505 | Cell Line | PANC1 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0506 | Cell Line | CFPAC1 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0507 | Cell Line | MIAPACA2 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0508 | Cell Line | CAPAN2 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0509 | Cell Line | BXPC3 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0510 | Cell Line | HS766T |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0511 | Cell Line | SU8686 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0512 | Cell Line | PSN1 |   | WWOX | S_not | NA | Not Known | 22.00% | Bisulphite Sequencin | 15073125 | ||||||||||||||
| PCM0541 | Cell Line | PANC1 |   | CEBPA | S_not | NA | Not Known | NA | Bisulphite Sequencin | 19035457 | ||||||||||||||
| PCM0542 | Cell Line | HPAC |   | CEBPA | S_not | NA | Not Known | NA | Bisulphite Sequencin | 19035457 | ||||||||||||||
| PCM0543 | Cell Line | MIAPACA | NA | CEBPA | S_not | NA | Not Known | NA | Bisulphite Sequencin | 19035457 | ||||||||||||||
| PCM0544 | Cell Line | MIAPACA2 |   | TMS1 | S_not | NA | High | NA | Bisulphite Sequencin | 21036703 | ||||||||||||||
| PCM0545 | Cell Line | ASPC1 |   | TMS1 | S_not | NA | Low | NA | Bisulphite Sequencin | 21036703 | ||||||||||||||
| PCM0546 | Cell Line | PANC1 |   | TMS1 | S_not | NA | Low | NA | Bisulphite Sequencin | 21036703 | ||||||||||||||
| PCM0547 | Cell Line | MIAPACA2 |   | SLC5A8 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 18437076 | ||||||||||||||
| PCM0548 | Cell Line | CAPAN2 |   | SLC5A8 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 18437076 | ||||||||||||||
| PCM0549 | Cell Line | PANC1 |   | SLC5A8 | S_not | NA | Not Known | NA | Bisulphite Sequencin | 18437076 | ||||||||||||||
| PCM01071 | Cell Line | CFPAC1 |   | TNFRSF10C | S_not | NA | Low | NA | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM01072 | Cell Line | PANC1 |   | TNFRSF10C | S_not | NA | High | NA | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM01073 | Cell Line | BXPC3 |   | TNFRSF10C | S_not | NA | High | NA | Bisulphite Sequencin | 21269942 | ||||||||||||||
| PCM01074 | Cell Line | SW1990 |   | TNFRSF10C | S_not | NA | High | NA | Bisulphite Sequencin | 21269942 | ||||||||||||||




|
||||||||||||||||||||||||
