Help Page
This page provides help to the users of AlgPred 2.0 on how to efficiently use the various prediction modules. For each module which is implemented in AlgPred 2.0, here we provide the figures and description on how to use that module.
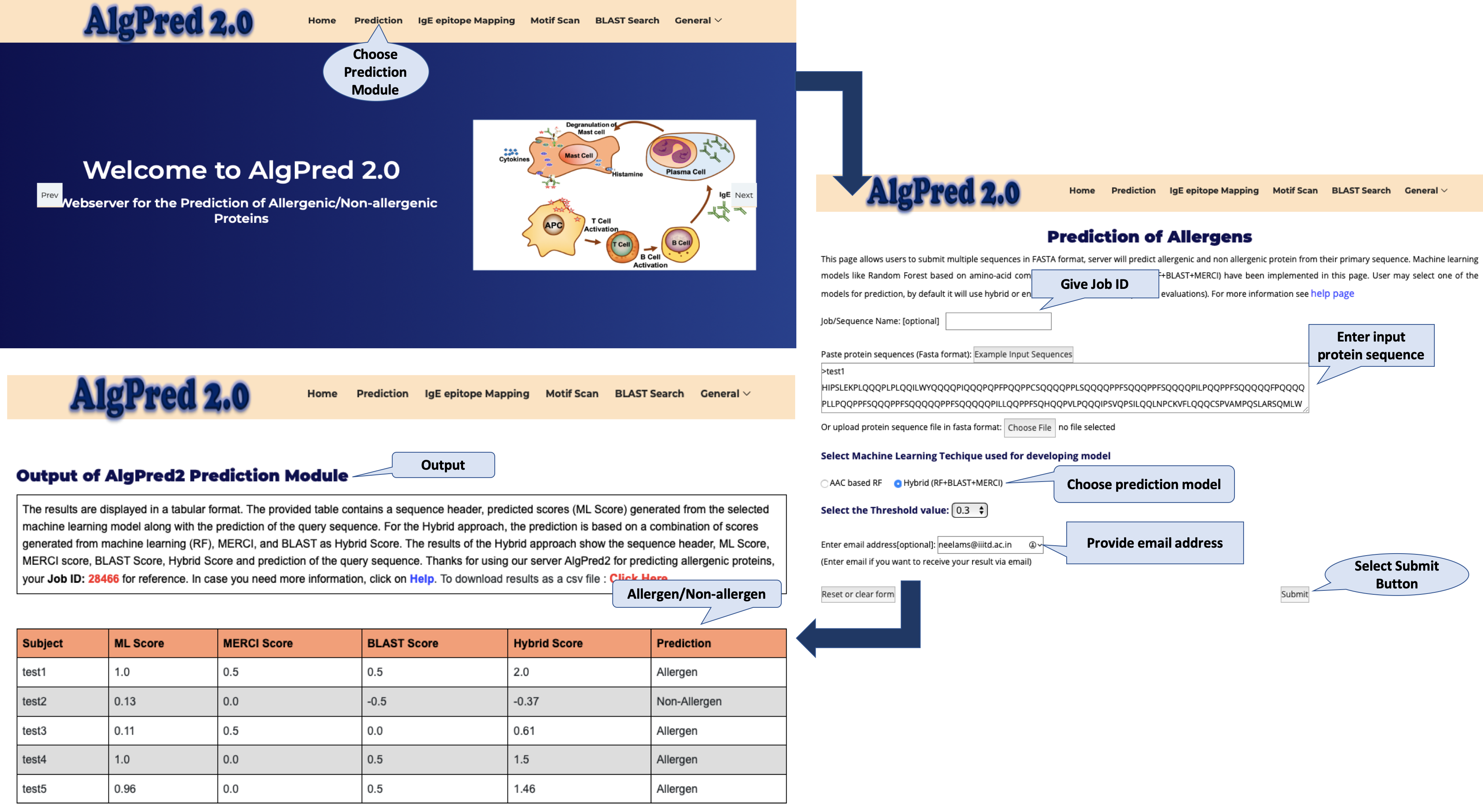
Prediction Module
This module predict allergenic/non-allergenic protein from its primary, i.e., amino acid sequence.
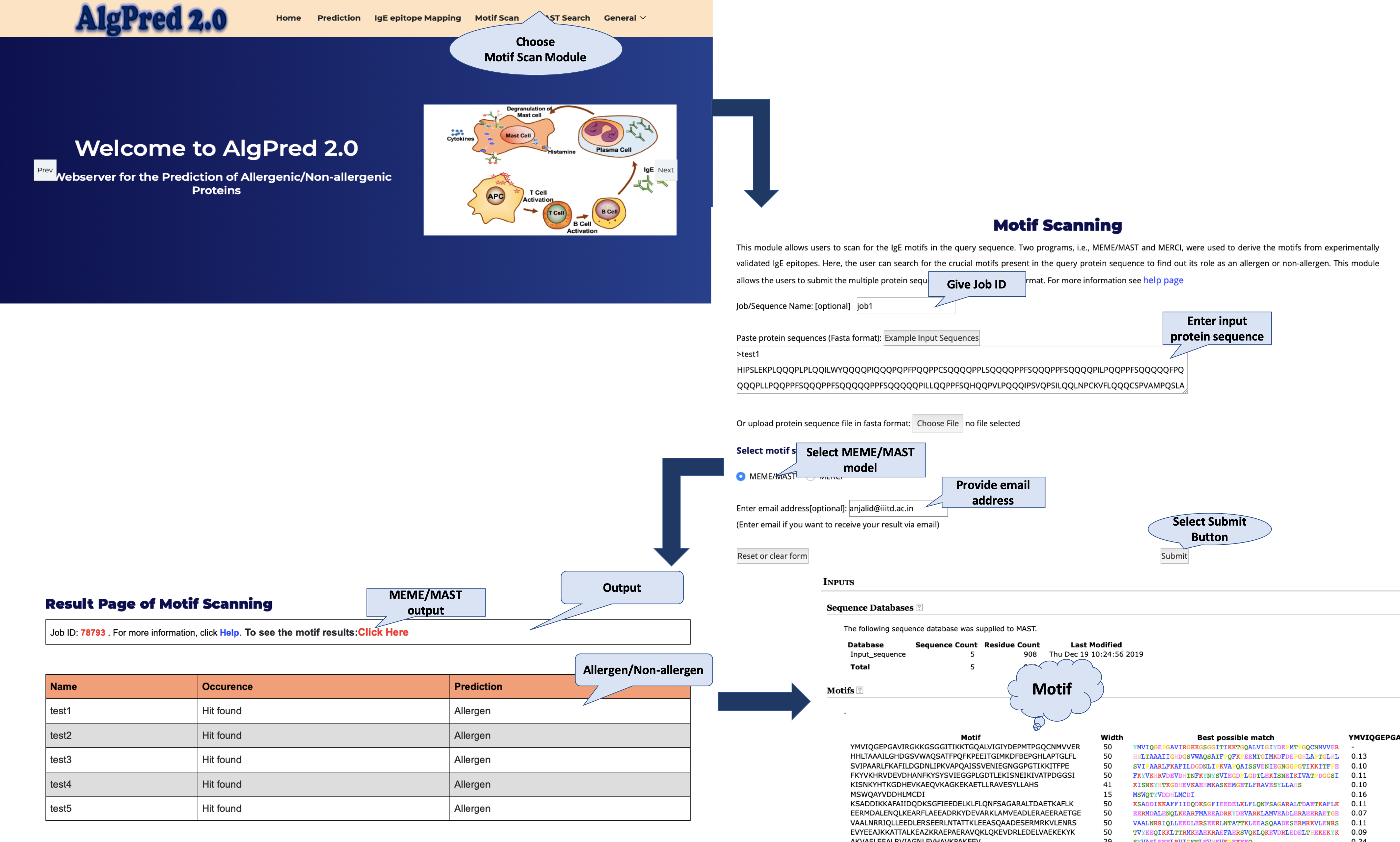
IgE epitope Mapping
This module is designed for mapping experimentally validated IgE epitopes on the query sequence.