Prediction |
| In the prediction menu user can submit a query, which may in form of peptides or proteins. In this menu user is provided with variable length or fixed length prediction for class specific B cell epitope prediction. |
Peptide prediction
|
| This section of webserver provides user a tool to predict peptide for its tendency of inducing particular class of antibody. This section has two different type of models, based on type of data used for model building: |
1: Variable length |
| The variable length page provides prediction of peptides with the machine-learning models based on variable length data set (BalanceVar) |
2: Fixed length |
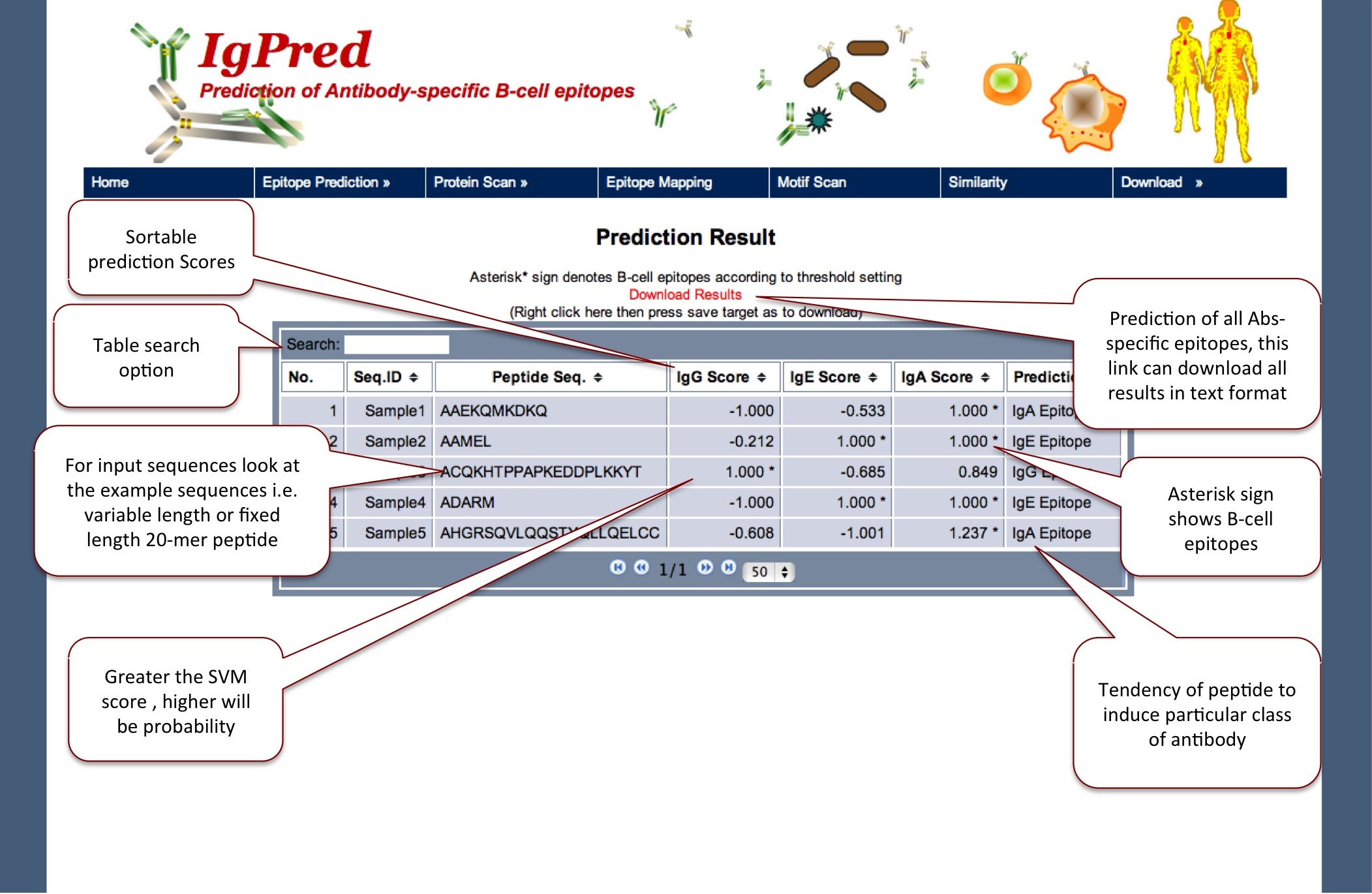
The Fixed length page provides prediction of peptides with the machine learning models based on fixed length data set (BalanceFix)
|
|
|
Protein prediction
|
| This tool assists users to identify antigenic regions on user defined protein sequences, which can induce particular class (IgG, IgE or IgA) of antibody and compare these regions visually.In this tool user can submit full length protein sequences |
1: Mapping with variable window |
| This prediction relies upon the machine learning model based on variable length data, so user can select any of the provided window from 4 to 20 and proceed for prediction. |
2: Mapping with Fixed window |
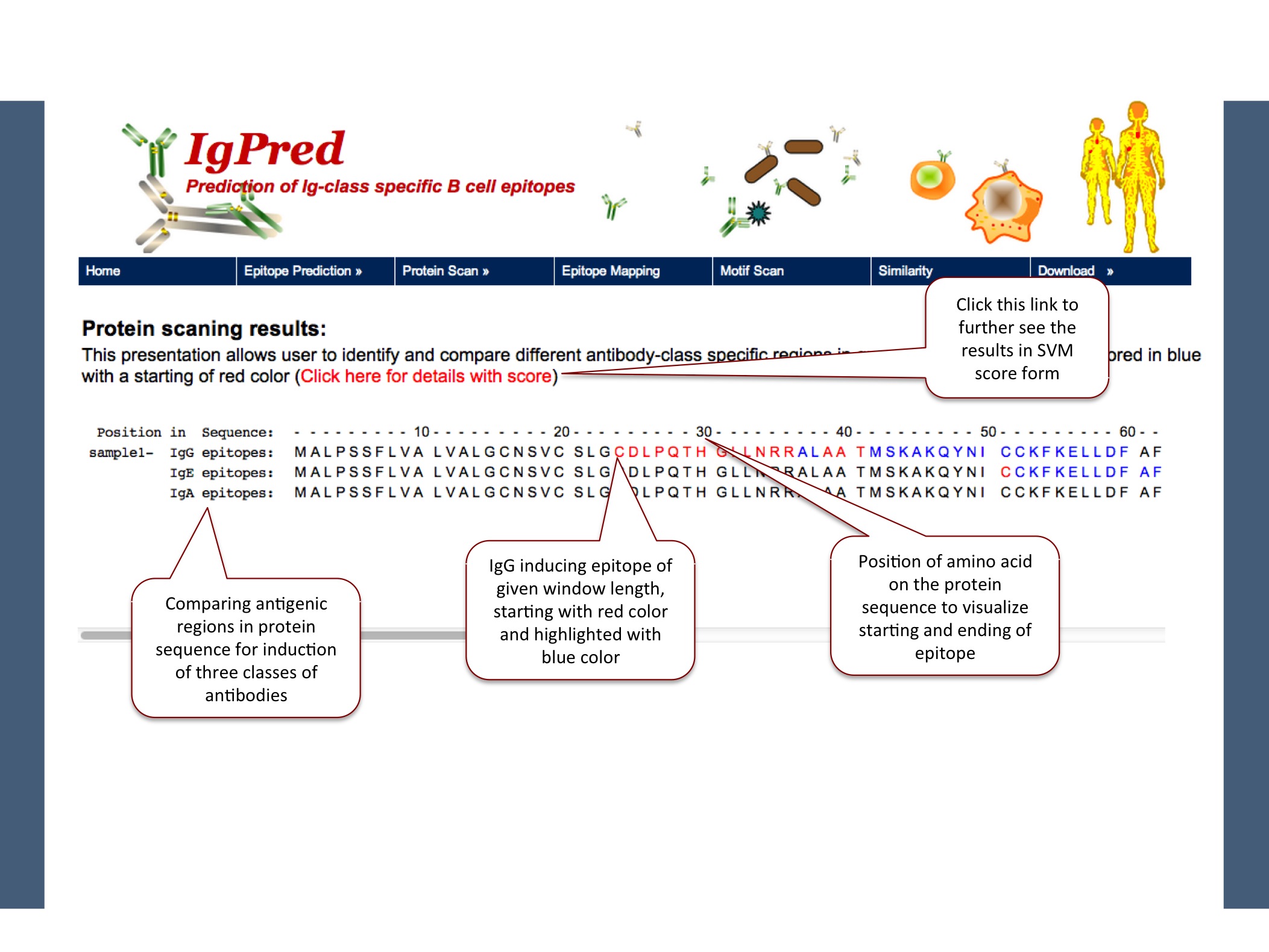
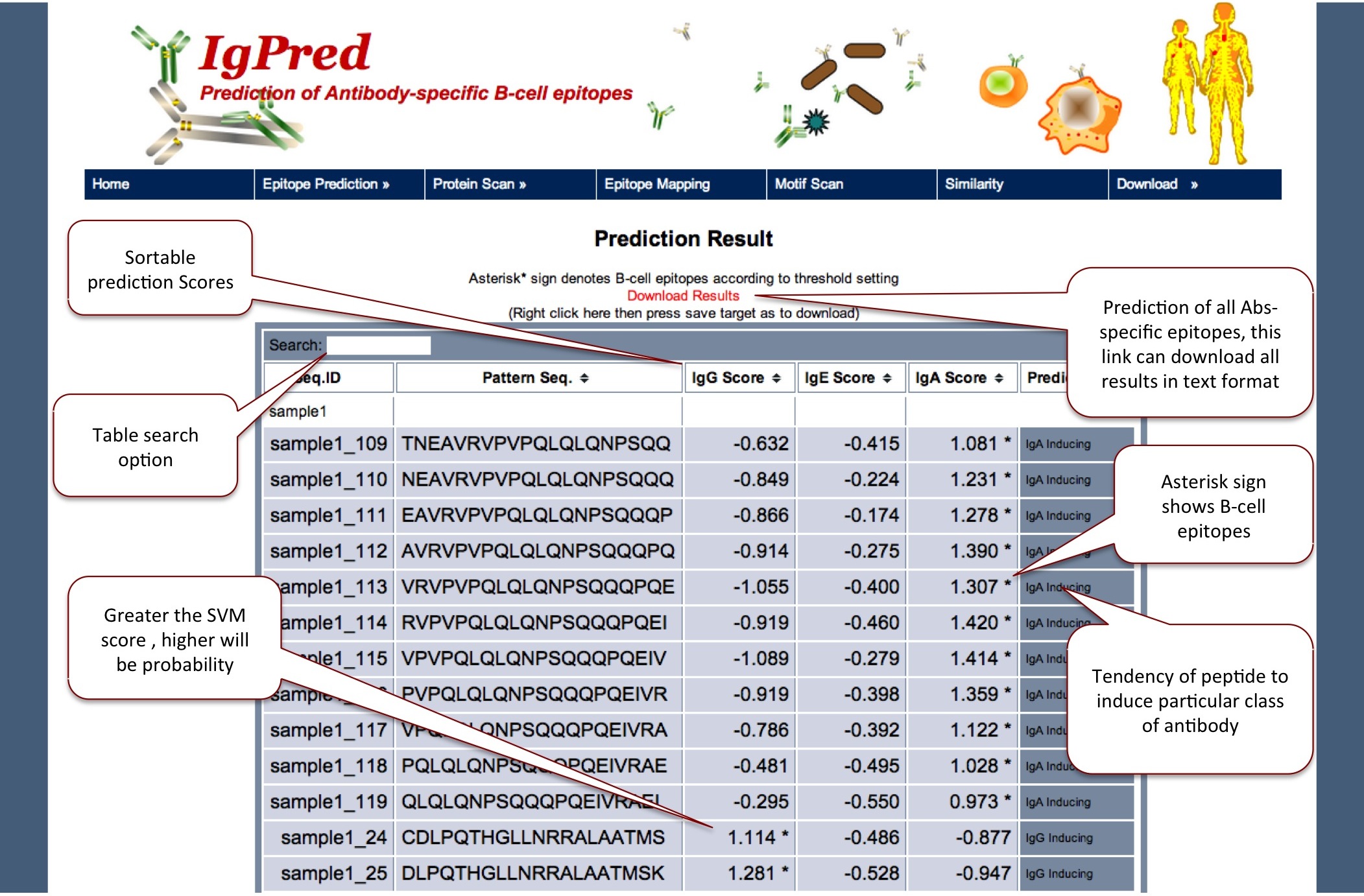
Here the models are based on fixed length dataset (20-mer), so user can give only the protein sequence and by default window will be taken as 20 for prediction.
|
|
|
|
Protein mapping with experimental data
|
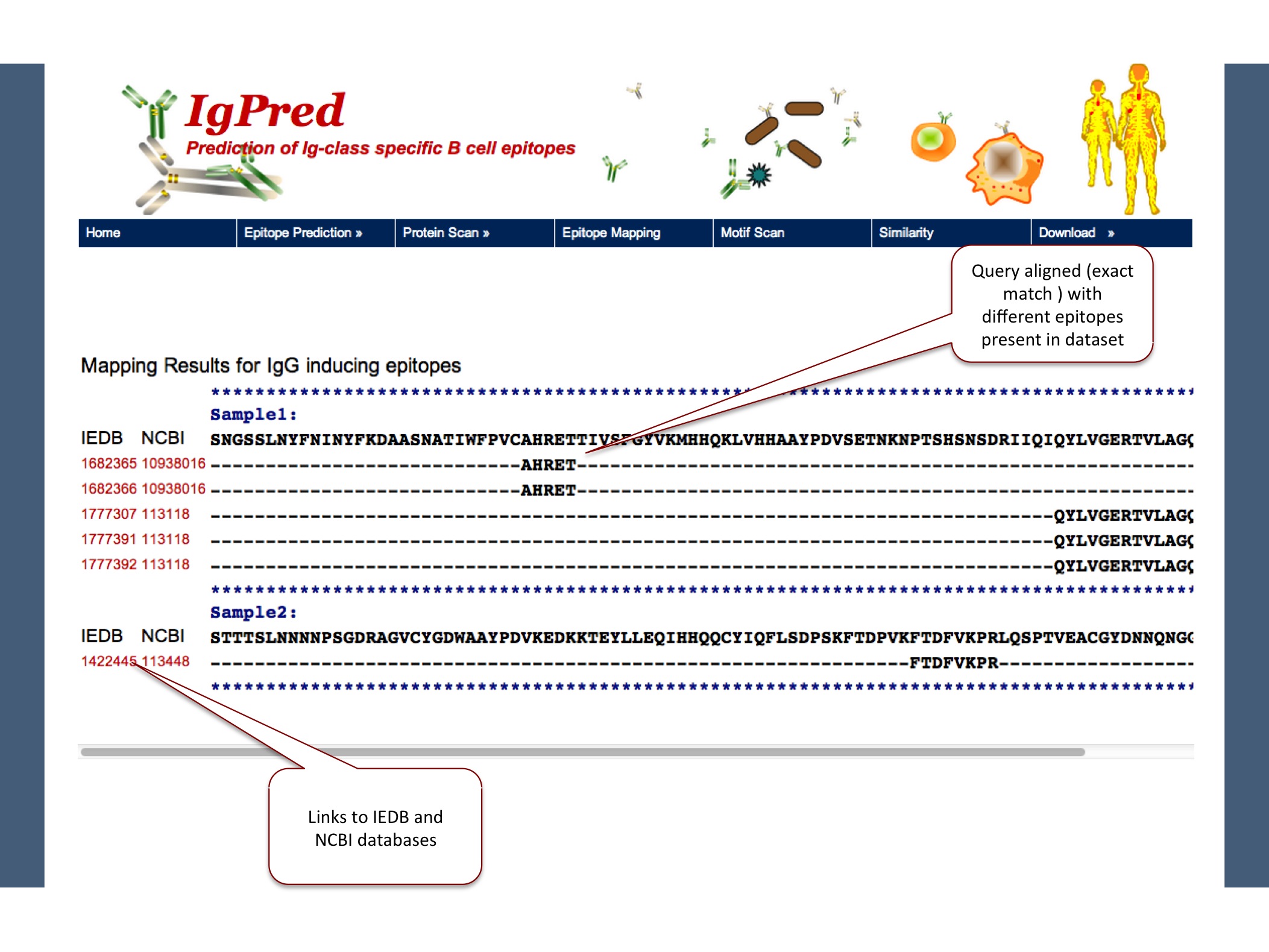
We have also provided an option of exact matching of query with our dataset, keeping antibody classes in mind. Here matches are also provided with link to IEDB along with NCBI accession number.
|
|
|
Motif Scan
|
To search the presence of Antibody-class specific motifs, discovered by MEME software, user can select the options of antibody class given in page. This tool gives the result in MAST default format where motifs can be visualized, aligned with the query sequence
|
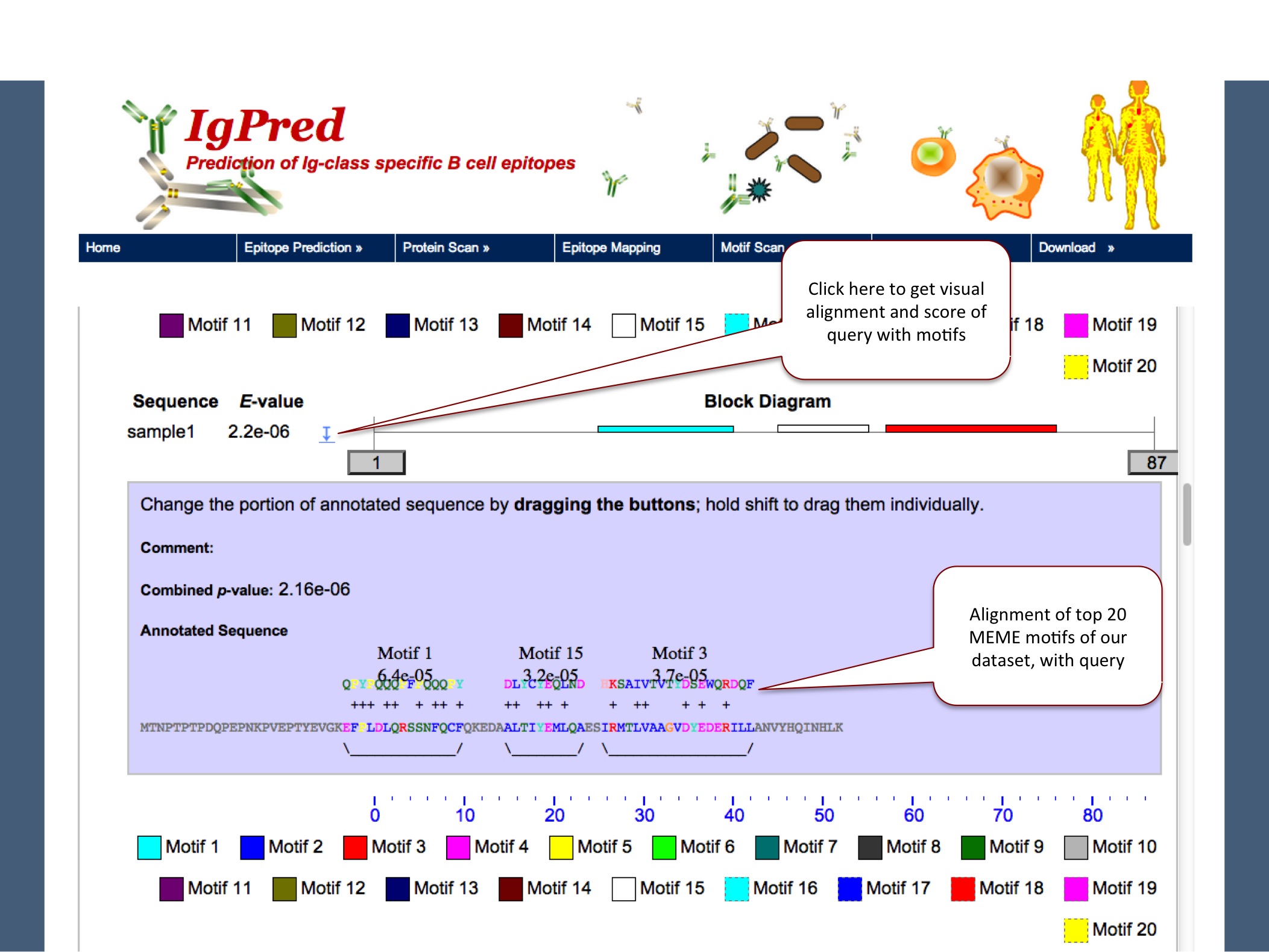
|
|
Smith-Waterman Similarity Search
|
This tool enables user to look at the similarity of query sequence with the epitopes present in our dataset. Result will provide the similar epitopes for which experiments have been done and reported in IEDB database, for induction of that particular class of antibody.
|
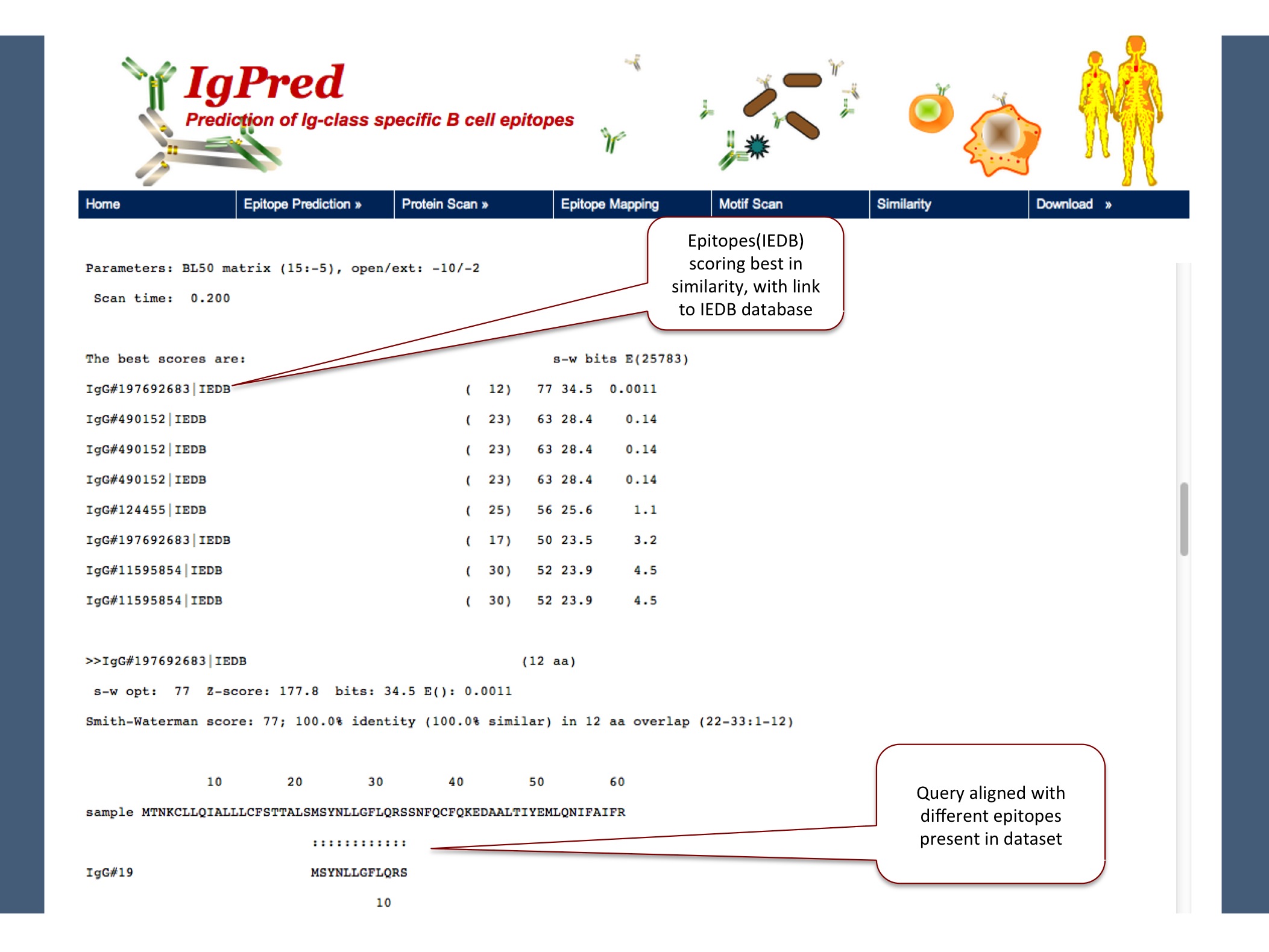
|
|
Algorithm
|
| This page explains the techniques and features, which have been used in our study of analysis and prediction. |
Download
|
| This link provides download options for every data set, which have been used in our study. |
| Supplementary Data: This page enables user to download the supplementary figures, tables, results and other data. |
| Data Set: Here user can get the main data set for all classes of epitopes. |
|
Frequently Asked Questions |
Q1. What is IgPred?
Ans. IgPred is a web server which provides user different tools such as motif match, similarity match, exact match of query with Antibody class-specific B cell epitope data and the prediction of query for what class of antibody is more probable to be generated with the query as antigen. etc.
Q2. What is the source of IgPred dataset?
Ans. IgPred dataset has been taken from IEDB database
Q3. What is motif scan? How does it work?
Ans. The motif scan tool is based on MEME-MAST program, where we have made use of MEME to discover top scoring motifs. Further the MAST program is used to search the query for the presence of motifs given by MEME. When a query is submitted, MAST gives motifs aligned with the query in the output.
Q4. How "Similarity" is different from "Mapping with experimental data"
Ans. We have given two different search options. First one is Similarity where similarity between query and our dataset will be looked for with the help of Smith Waterman Algorithm, while "Mapping with experimental data" enables user to see exact match of query with our dataset
Q4. How is Peptide prediction different from Protein Prediction ?
Ans. In Peptide prediction, the peptide is taken as a whole and we do not provide any window selection, while in Protein prediction we give option of selecting window length for the cutting of protein in to peptides. Furthermore, variable window options in protein module have been given for those who want length of peptide from the query protein, between 4-20 and similarly length is fixed at 20 in case of fixed window.
|





