Welcome to Help Page
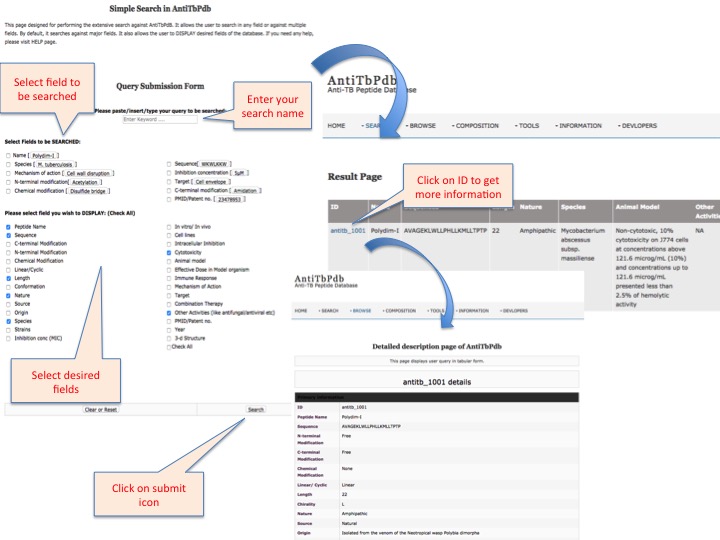
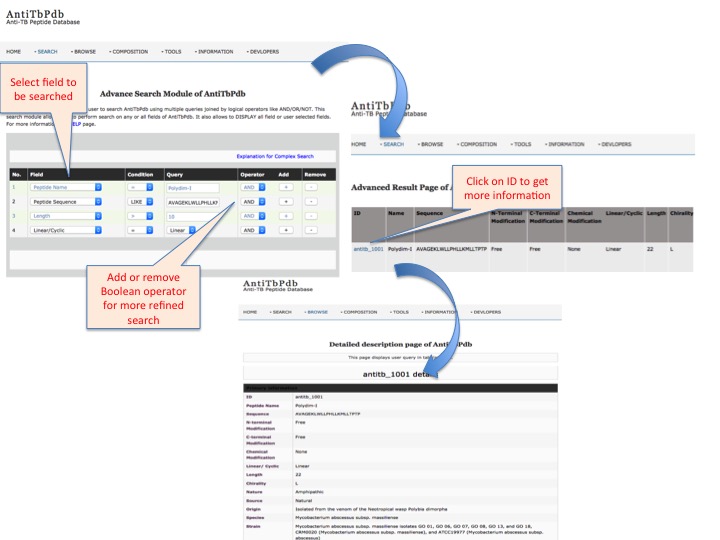
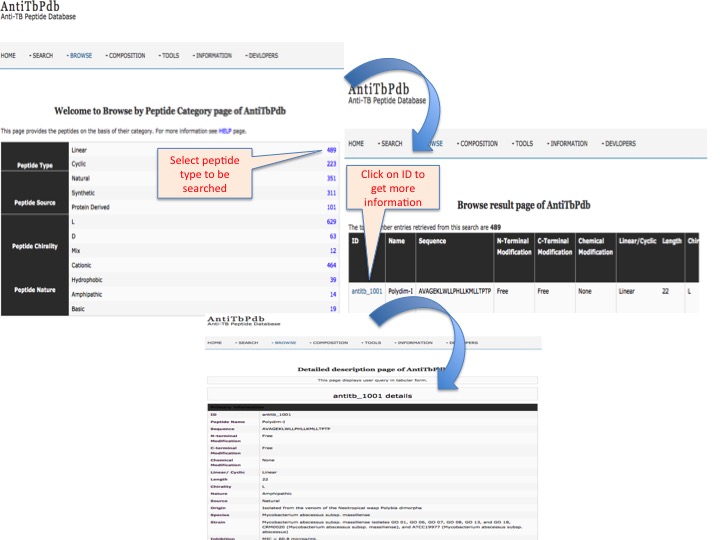
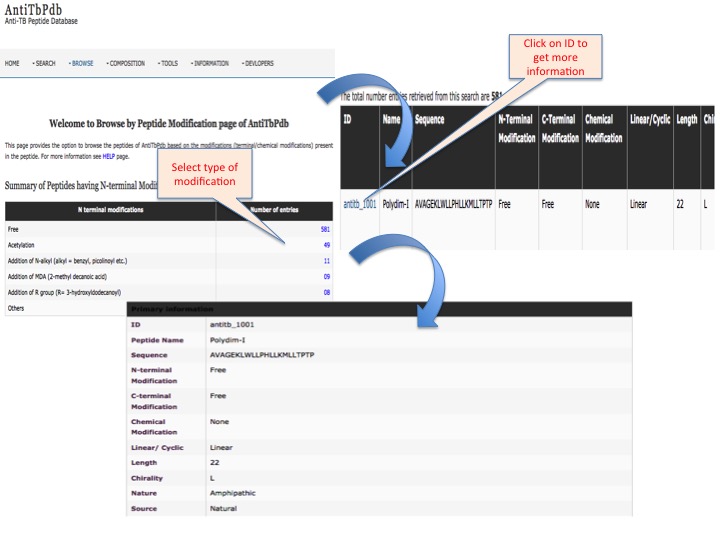
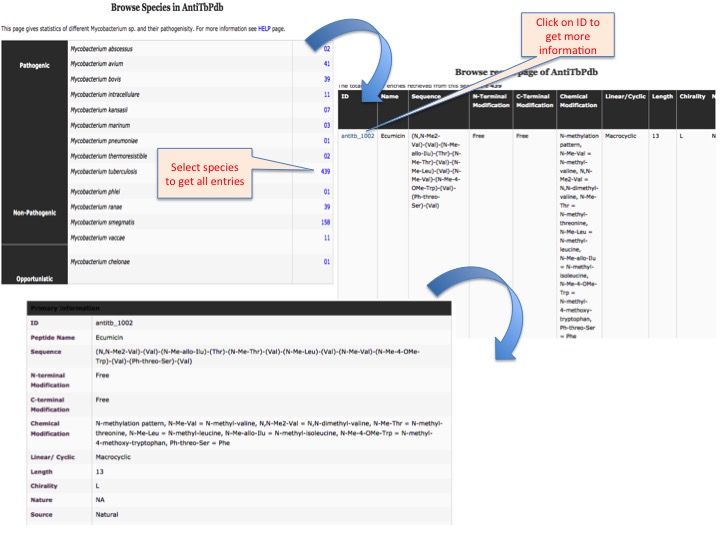
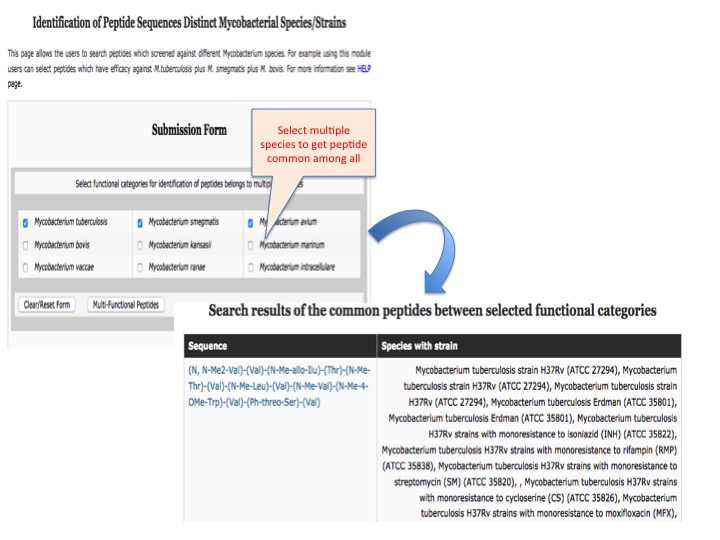
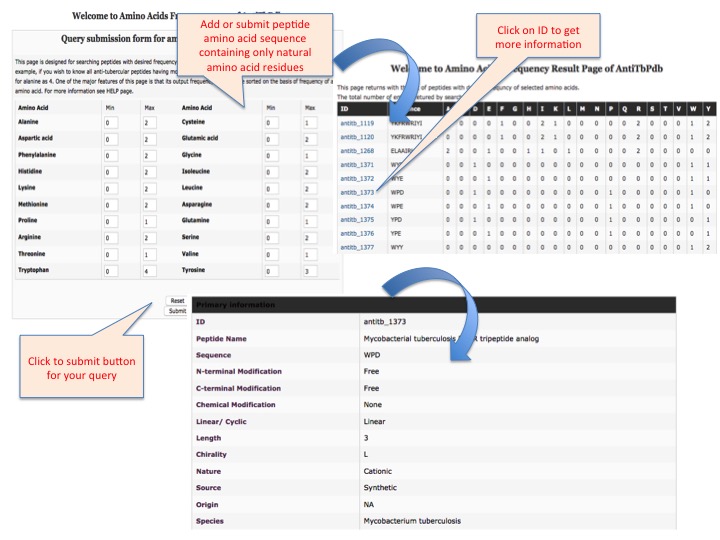
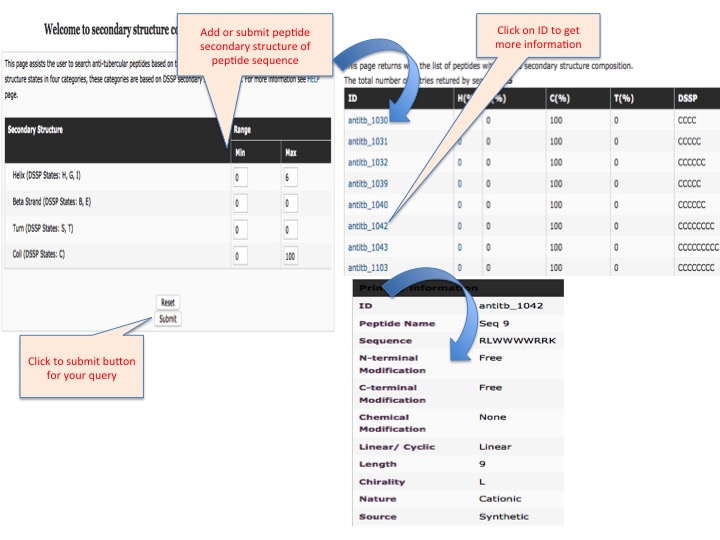
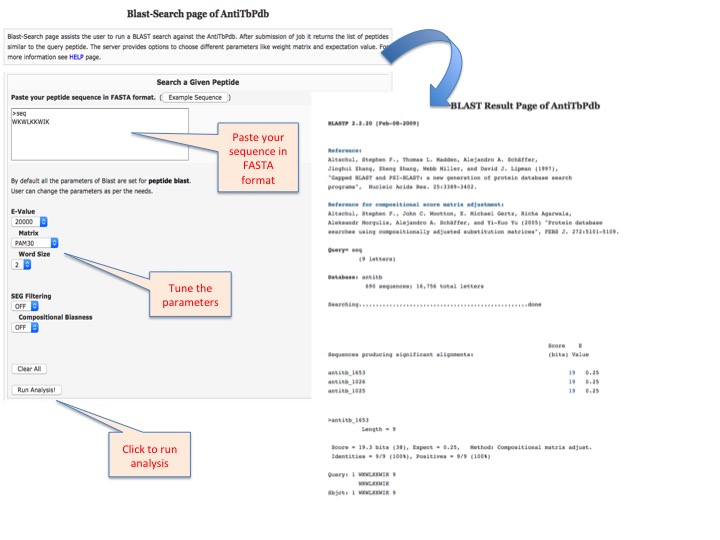
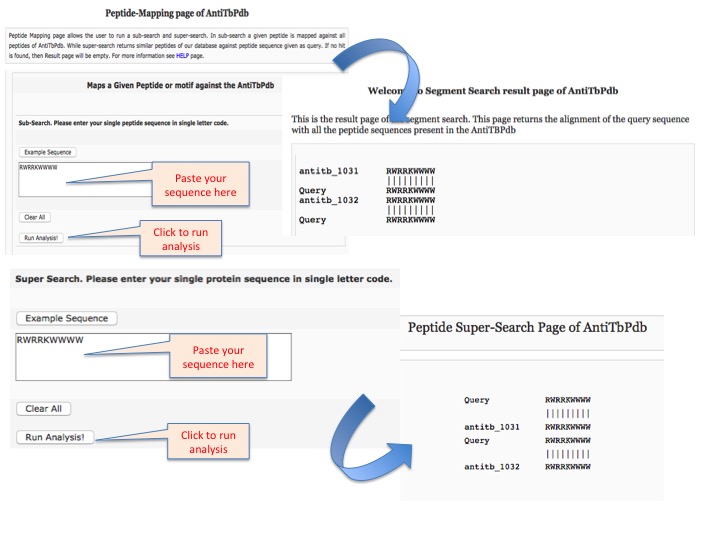
This page provides help to the users of AntiTbPdb database on how to efficiently use this repository of antitubercular peptide. For each module which is implemented in AntiTbPdb, here we provide the figures and description on how to use that module. Four general modules are provided with the respective.
Frequently Asked Questions (FAQs) |
|
Q1. What is AntiTbPdb? Ans. AntiTbPdb is an comprehensive manually curated repository of experimentally verified anti-tubercular or anti-mycobacterial peptides. Q2. Why AntiTbPdb? Ans. It comprising peptide entries from the literature available on Pubmed and patents. It will give all the information about the peptide associated peptides, like its sequence, modification, source, half life, length etc.This database will helpfull to identify and design antitubercular peptides. Q3. How to search into AntiTbPdb? Ans. A user can perform sequence or secondary structure-based searching to identify identical/similar entries present in all the peptide databases integrated in AntiTbPdb. |