Help Page of TumorHoPe |
| This page provides help on TumorHoPe database. |
SEARCH | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
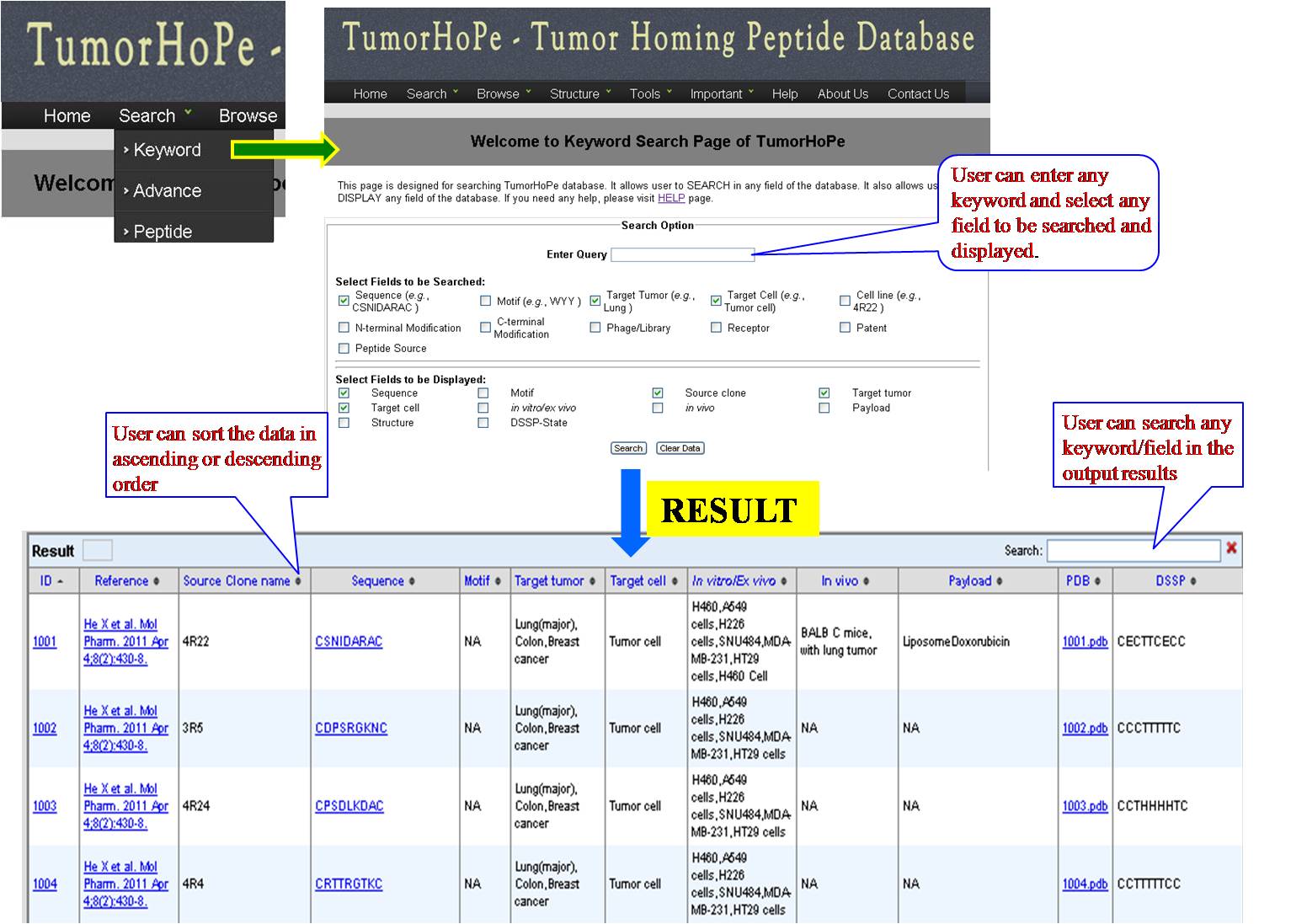
KEYWORD SEARCH This is a very simple and easy search option. User can search with single word for sequence (e.g. CSNIDARAC), motif (e.g. WYY), target tumor (e.g. Breast cancer), target cell (e.g. Endothelial cell), cell lines (e.g. MDA-MB) used in the experiments. Results will be displayed according to the selected fields to be dispalyed. e.g. sequence, research article, PMID etc. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
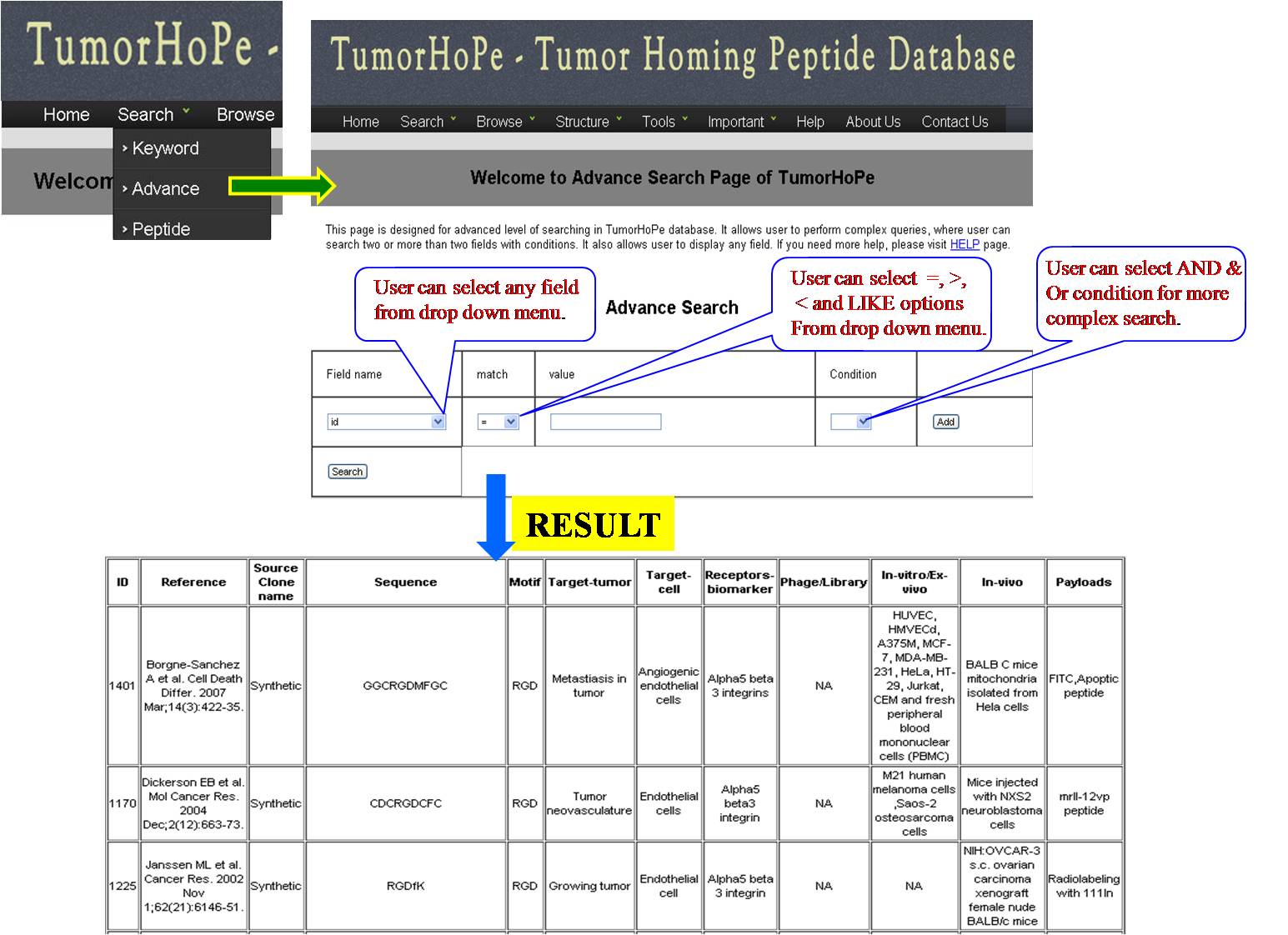
ADVANCE SEARCH This search has multiple options which give very refined result for the user. Here user has been provided with different search options Field name Match option (equal =, less <, greater > ). Value column is provided to add input from user e.g. ID, year, tumor related word etc. and then user can apply conditions(AND & OR). 1. If user is giving MOTIF in the field name and to match it with RGD, the user will opt for (equal=), result page will display all the peptides containing motif RGD. 2. If condition AND is applied then another search field will be opened, user can put value like lung, then results with motif RGD containing lung in the field selected will be displayed. 3. If condition OR with the above input, the user will get all peptides containing RGD motif and all of them will have word lung. 4. Match option LIKE, if user is opting for LIKE option then value must be put in between two % symbols e.g. % blood%. Here user will get result searched for blood in the given field option, if the user is putting % in front then result will be displayed where the word blood is in the beginning and if the % is put after word like blood %, then user will get result where blood word is in the end of the field which is choosen. *Note: While submitting, the advanced search for various conditons, the last search box in the conditions must be blank. By pressing search, desired result will be displayed. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
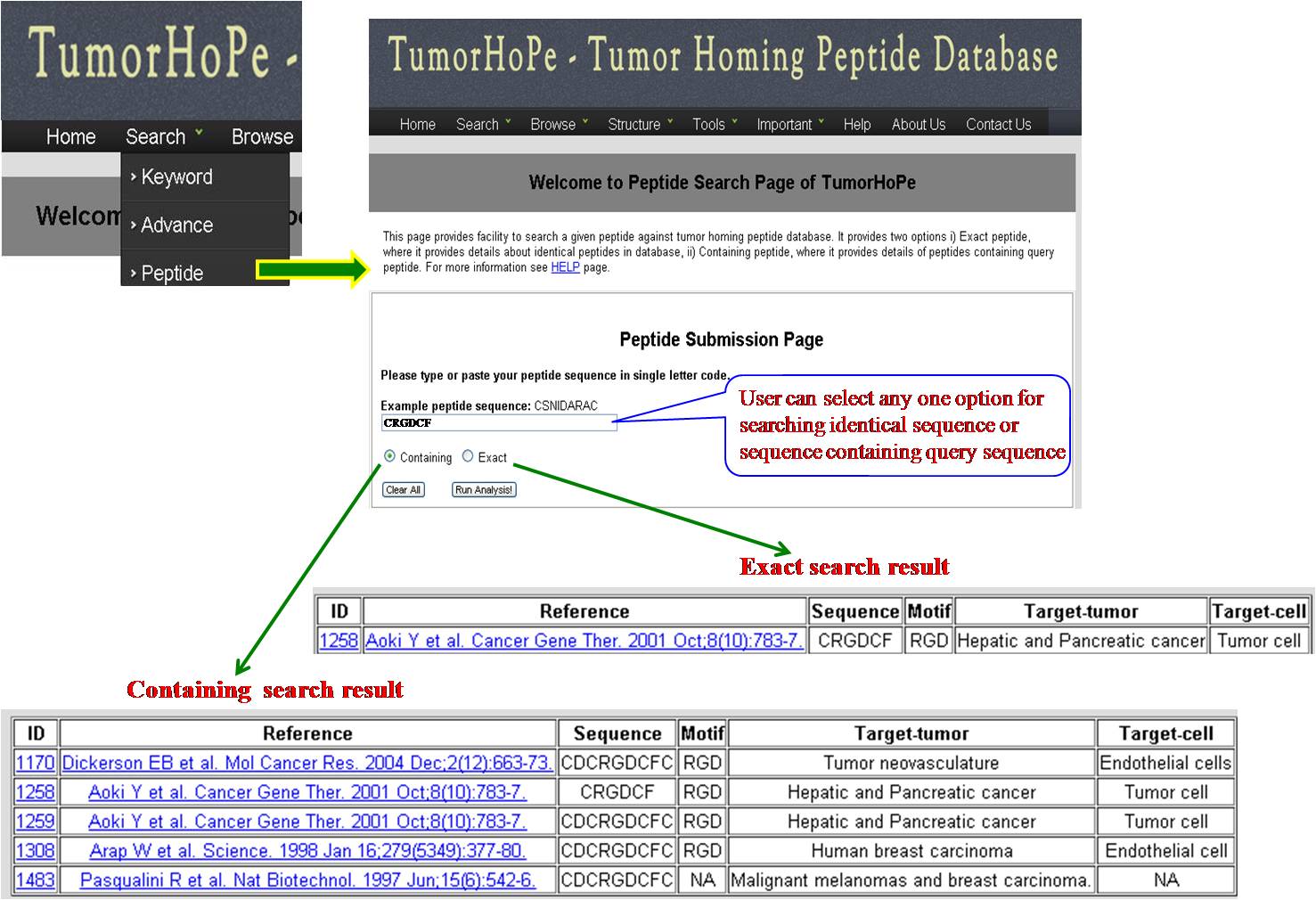
PEPTIDE SEARCH User can search the query sequence against whole database. Search result will provide user either with exactly identical peptide or matching peptides with the given query sequence. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
BROWSE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
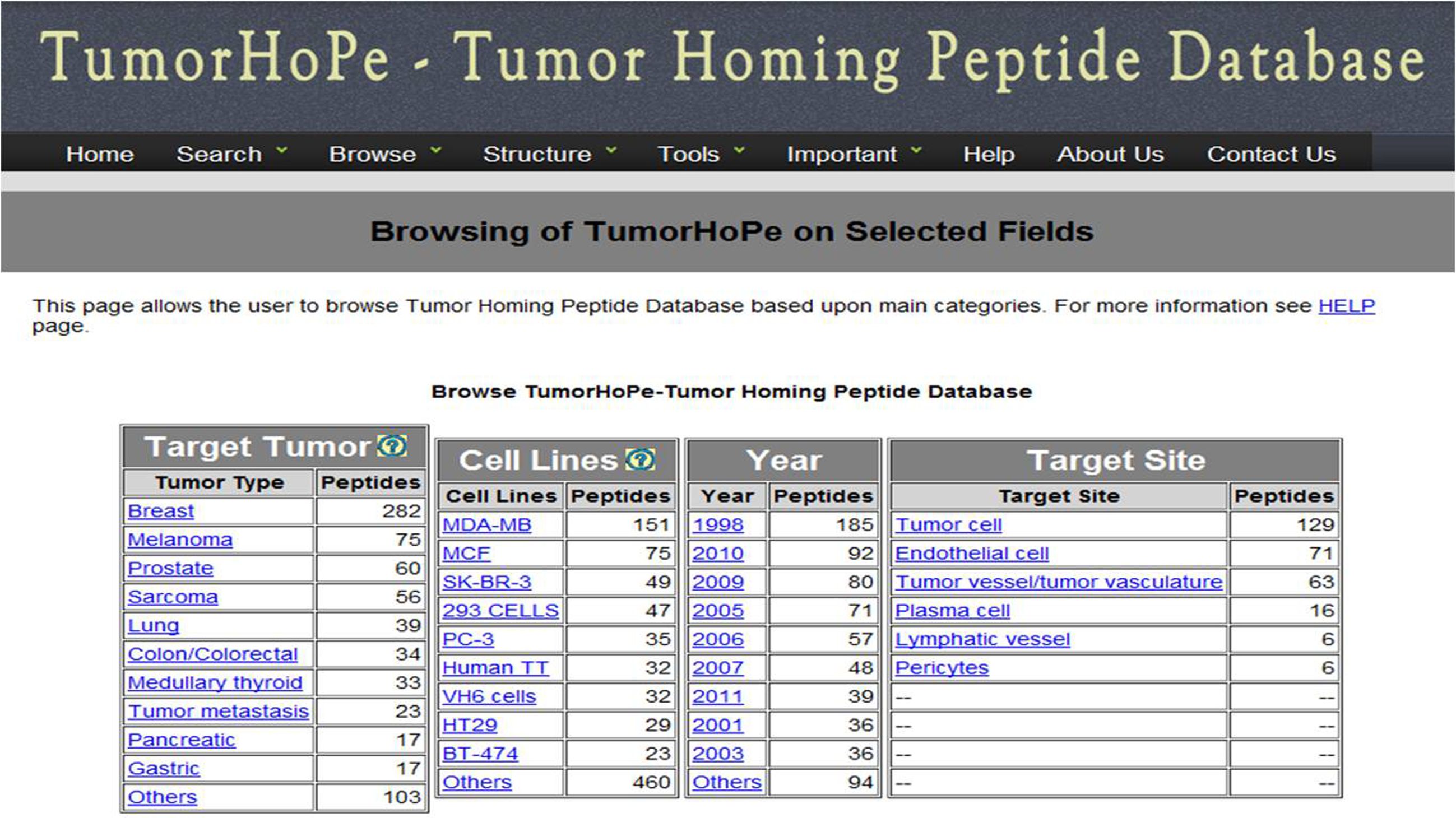
MAJOR FIELDS This interface allows user to browse database on the following four major fields (1) Target tumor (2) Cell line (3) Year of publication (4) Target site. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
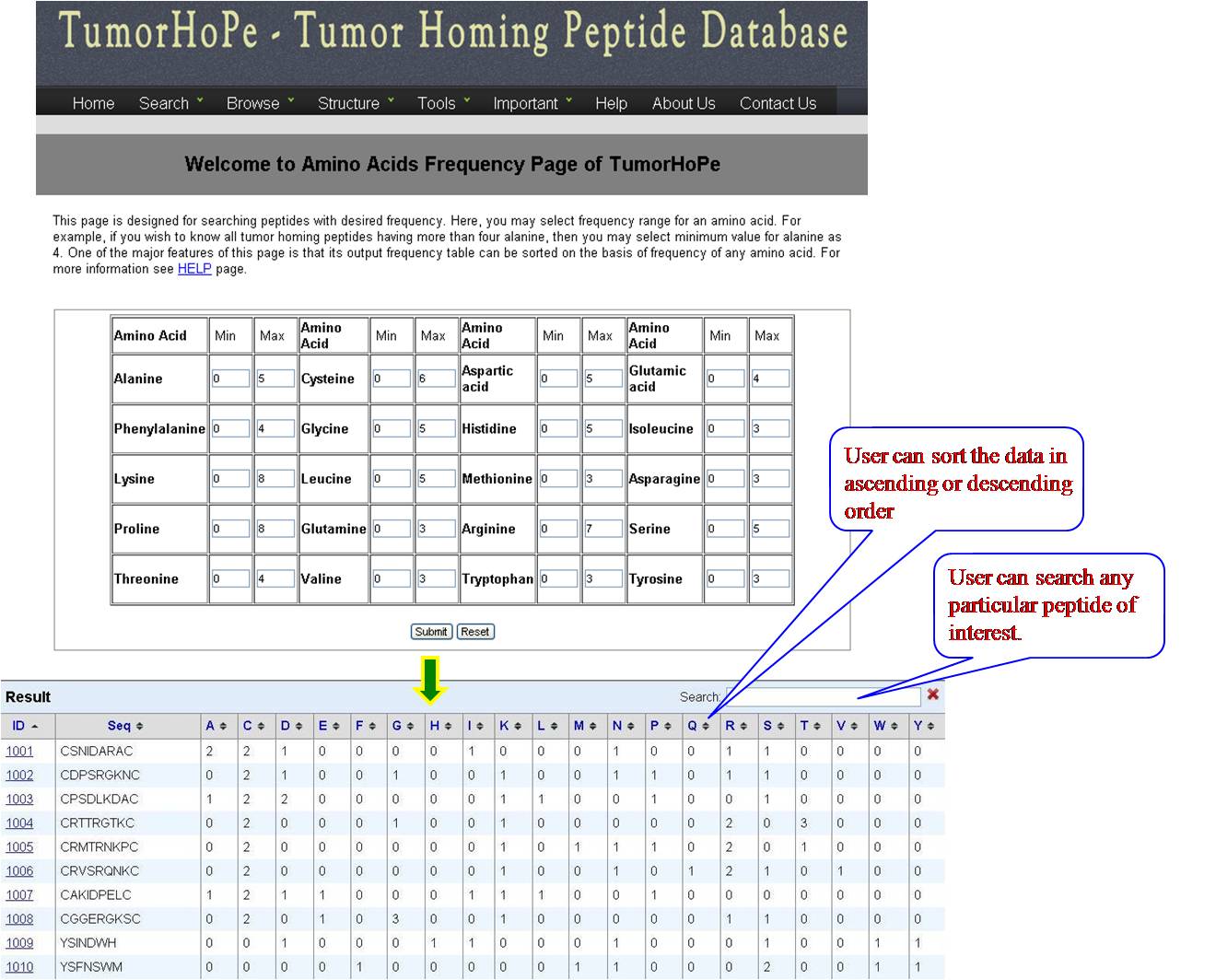
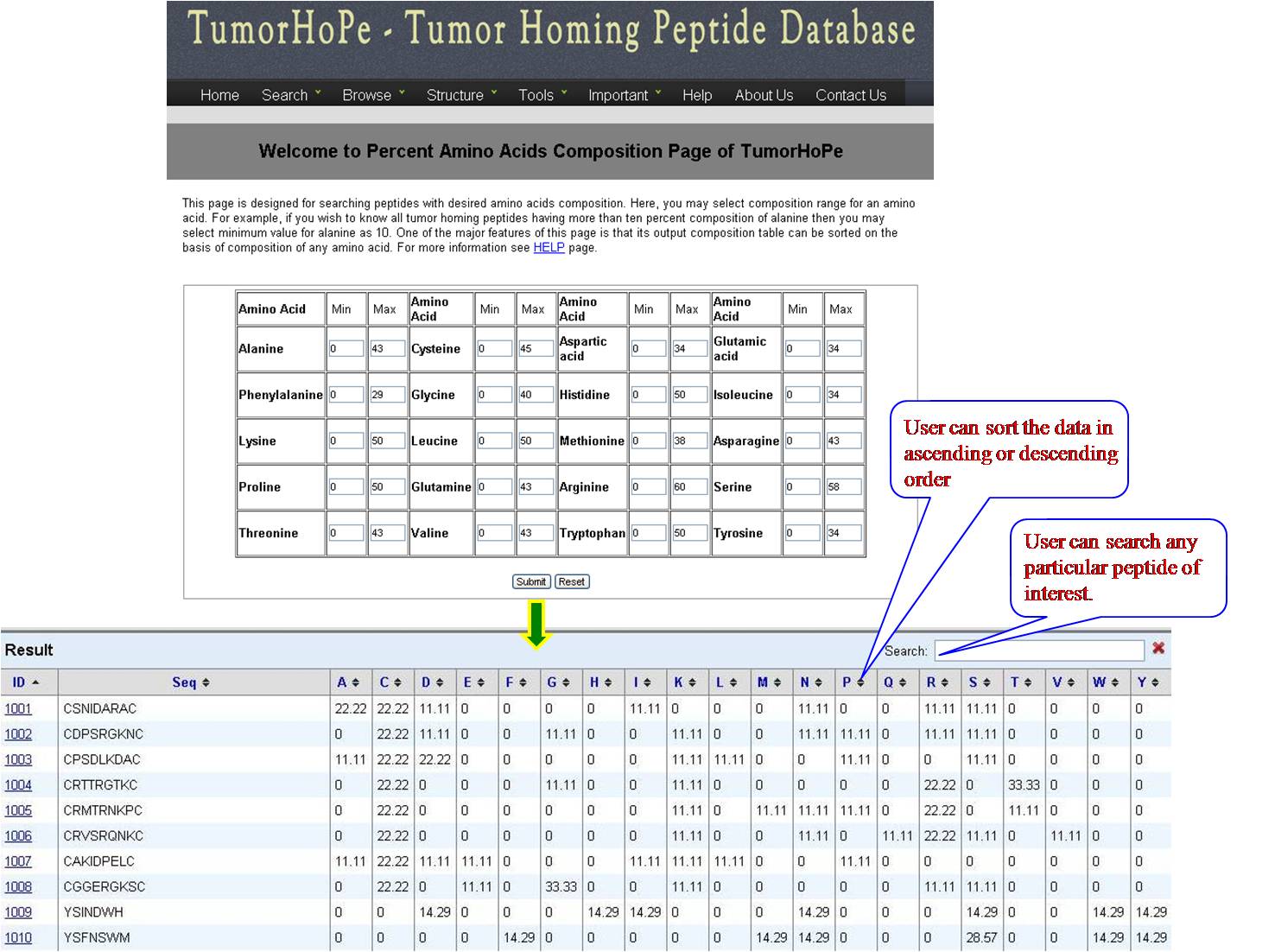
AA FREQUENCY AND AA COMPOSITION User can access the database for the amino acid frequency with range between (0-100) e.g. if the user enters lysine value 3 for minimum and 23 for maximum, the user will get list of all the sequences in the database with lysine value between 3 and 23. If the user is putting values for another amino acid along with lysine, e.g. Glycine 4 min and 18 max, then user will get list of sequences which has been sorted for lysine with desired value of glycine. Same procedure is followed for amino acid composition, Physical property frequency and composition. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
  | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
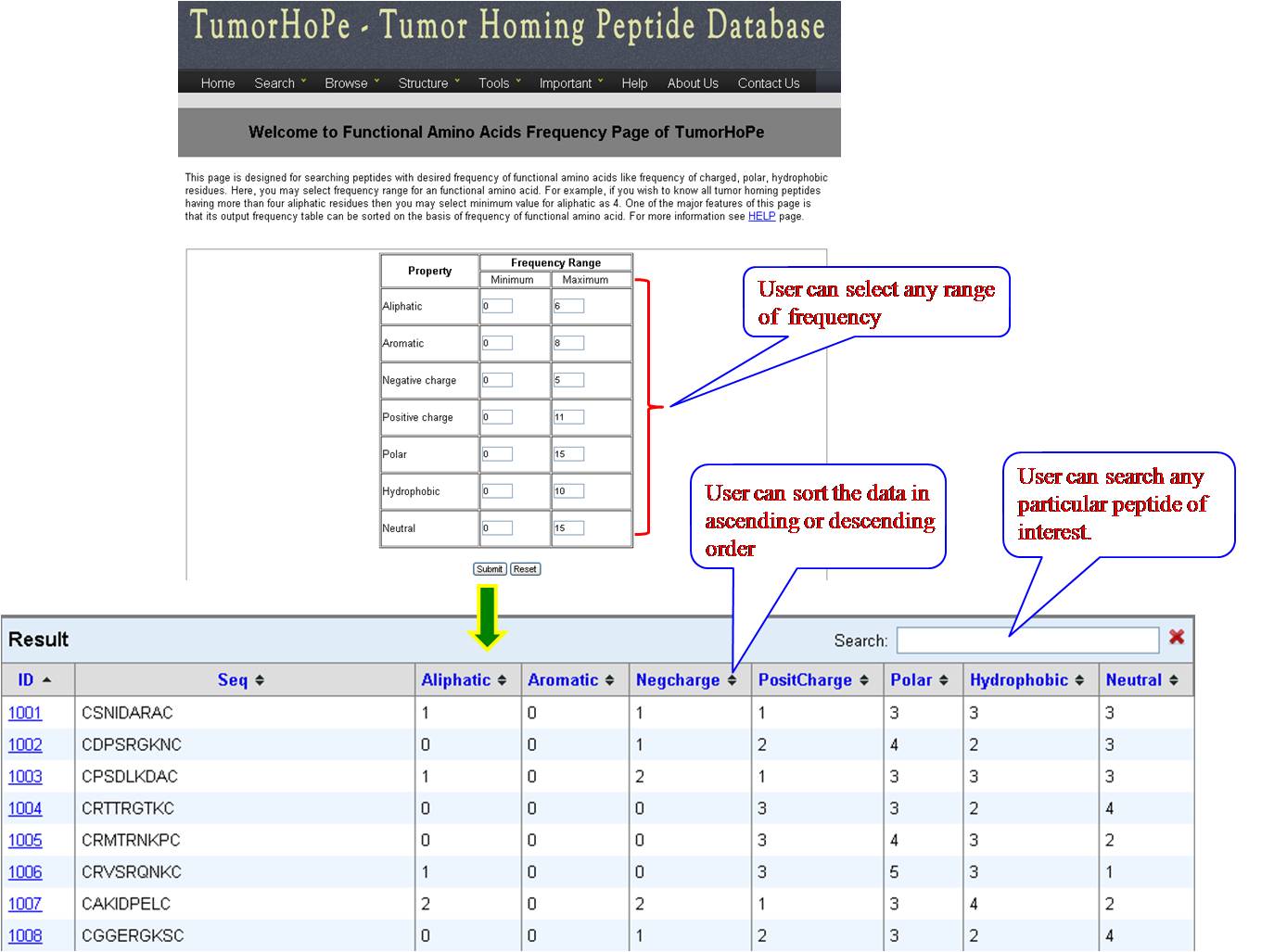
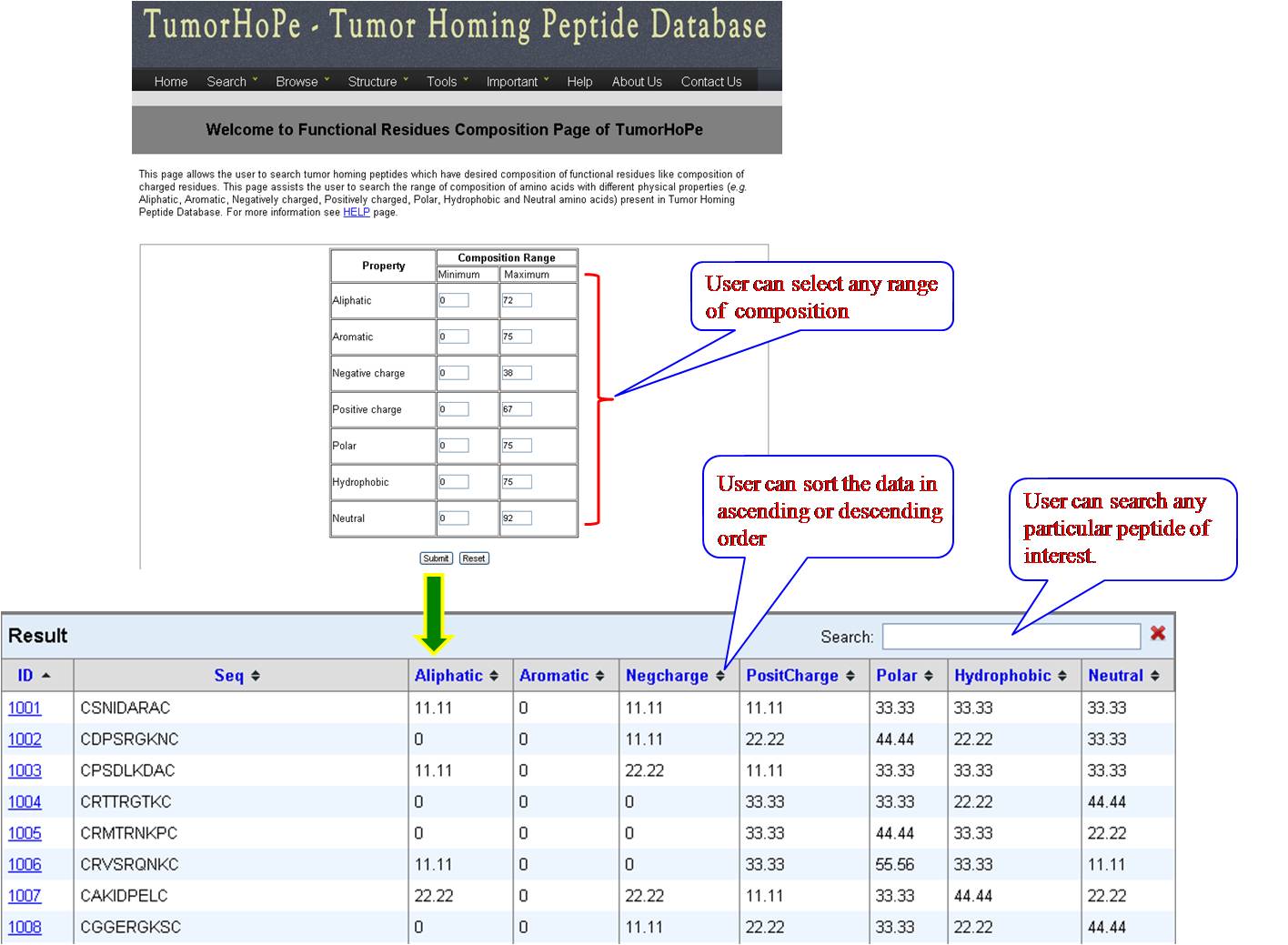
PHYSICOCHEMICAL PROPERTY (PP) FREQUENCY AND COMPOSITION User can extract tumor homing peptides that have desired frequency of certain types of residues like, positive charge, negative charge, polar residues. By default we have provided minimum and maximum values for our dataset. User can set the desired range for that property and peptides will be sorted e.g. if we give a range for aromatic property between (2-8), all peptides between this range will be listed in the output table. User can give combination of properties like aromatic property and positive charge, within the desired range. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
  | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
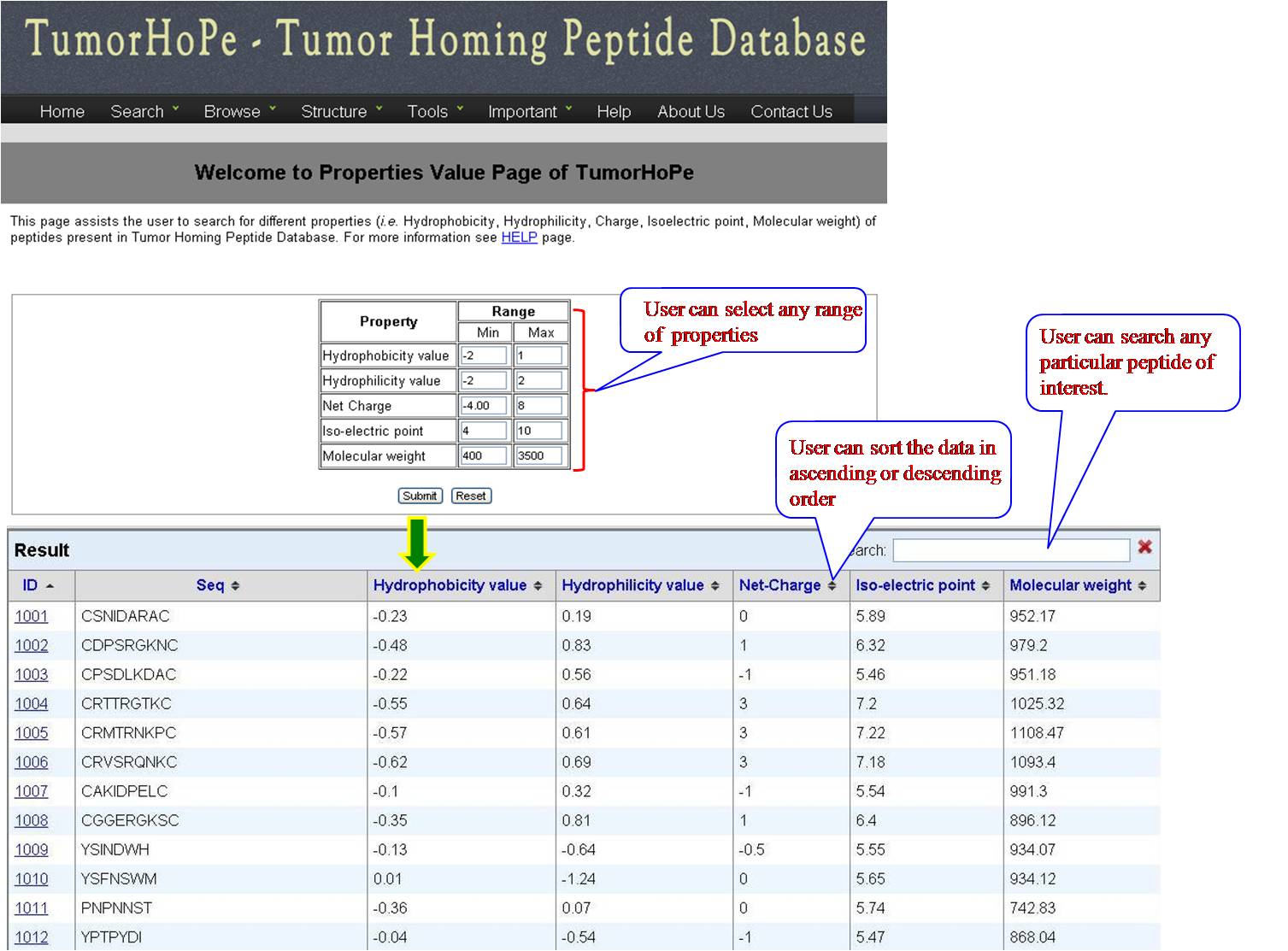
PHYSICOCHEMICAL PROPERTY VALUES This option provides user with list of tumor homing peptides with specified range (according to user) for Hydrophobicity value, Hydrophilicity value, Net charge, Isoelectric point and molecular weight. User has to fill the minimum and maximum value, and will get desired peptides in the above category. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
STRUCTURE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
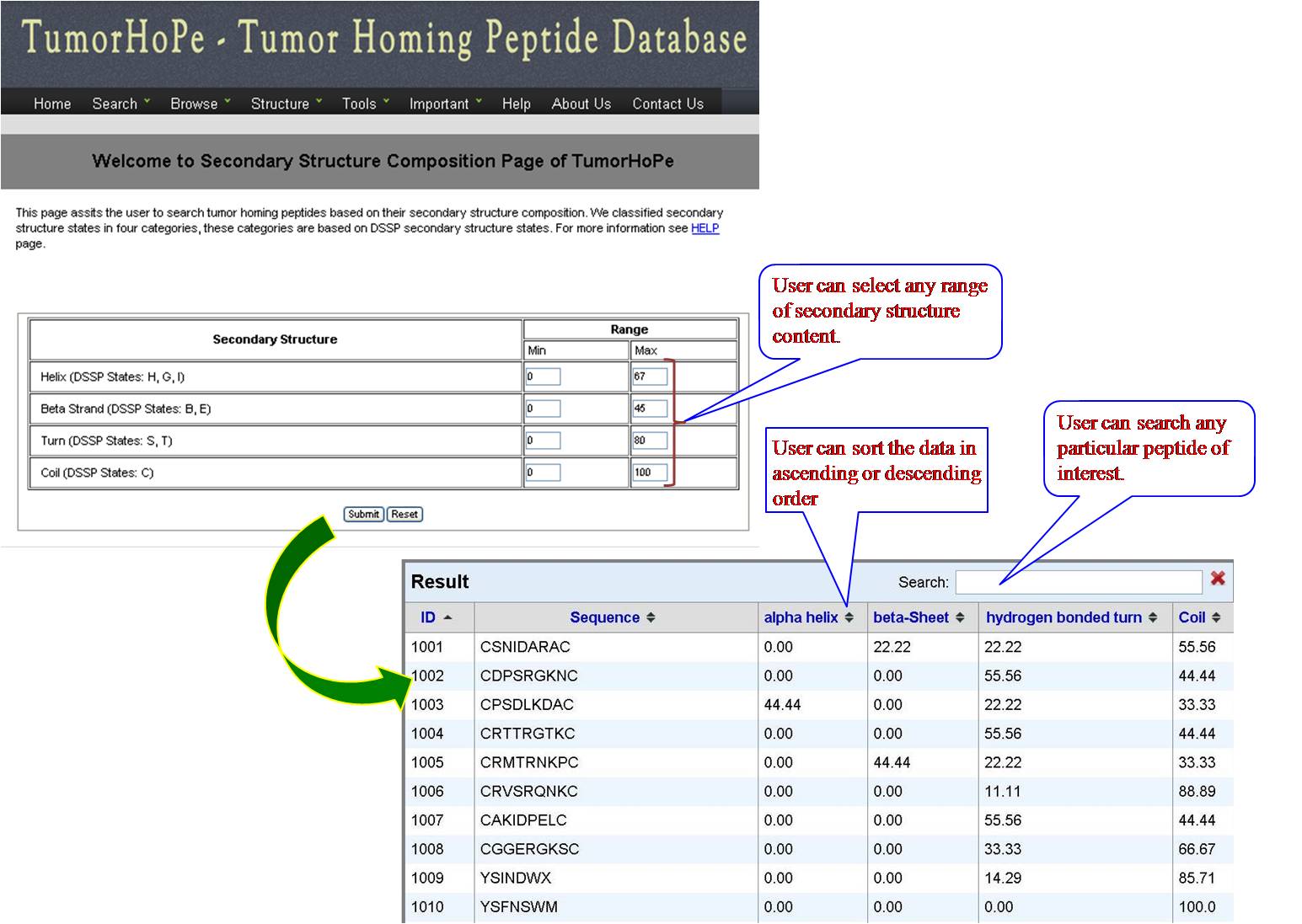
SS COMPOSITION User can get list of tumor homing peptides on the basis of percent composition of four different secondary structural states (Helix -H, Betasheet-E, Turn-T, Coil -C) with their desired range of minimum and maximum value. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
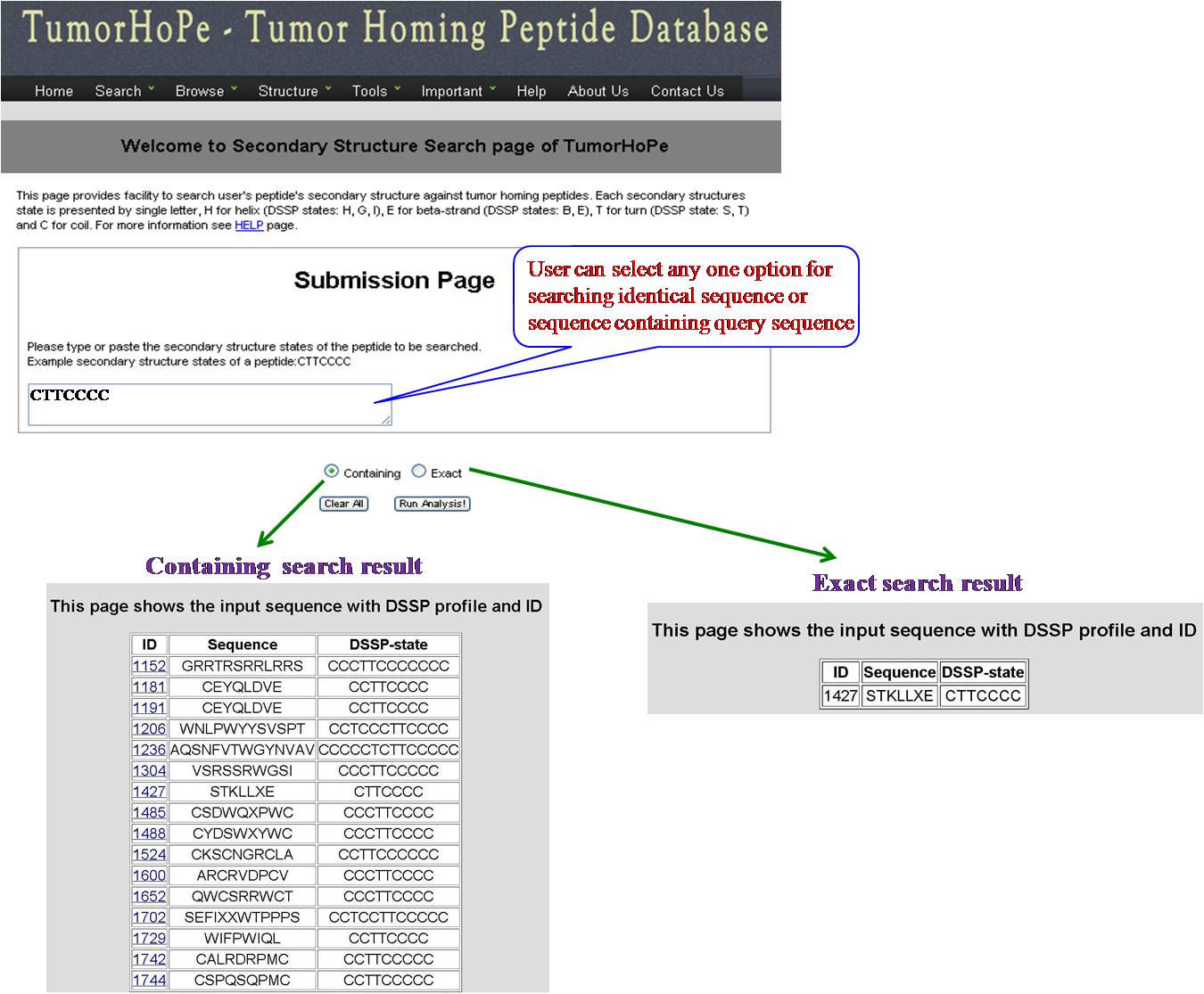
SS SEARCH In this option user has to submit the peptide sequence from the database and user will get predicted structural state. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
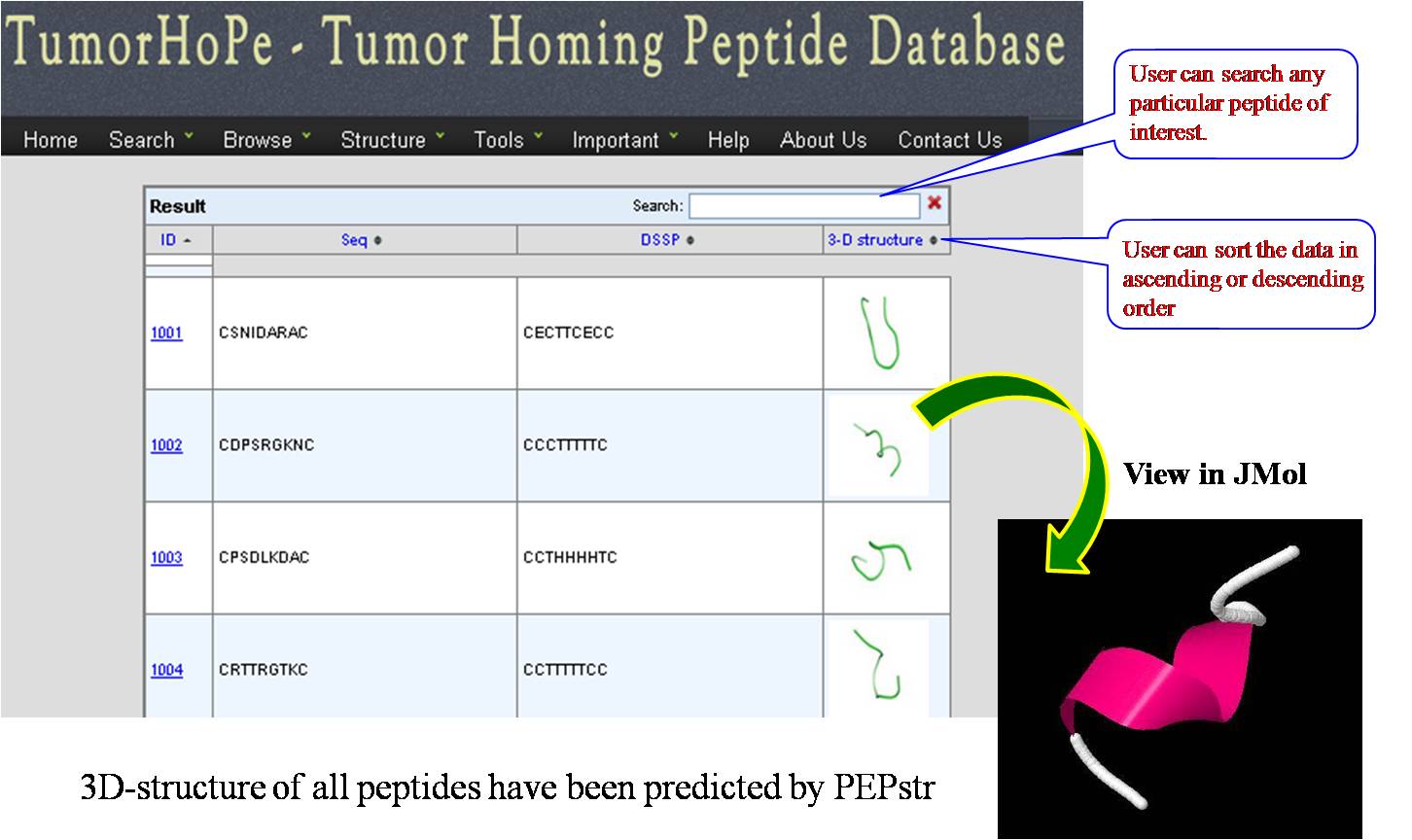
3D STRUCTURES 3D structures of all tumor homing peptides have been predicted using PEPstr. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
TOOLS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
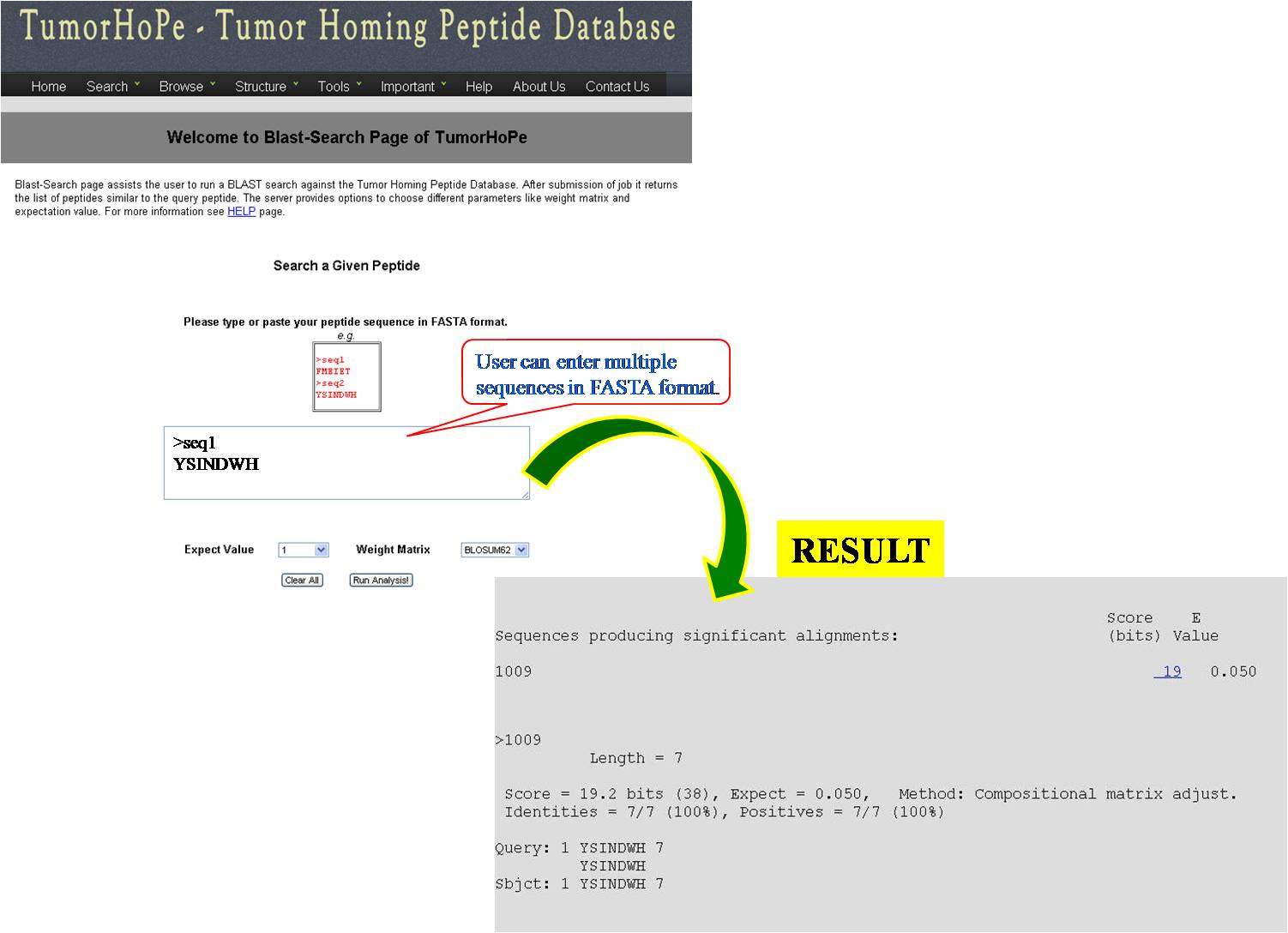
BLAST It helps the user to run BLAST against Tumor homing peptide database. This is similarity based search of any query sequence with those present in tumor homing peptide database. User can submit query sequence in single letter code in search field, it will display all tumor homing peptides similar to query sequence. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
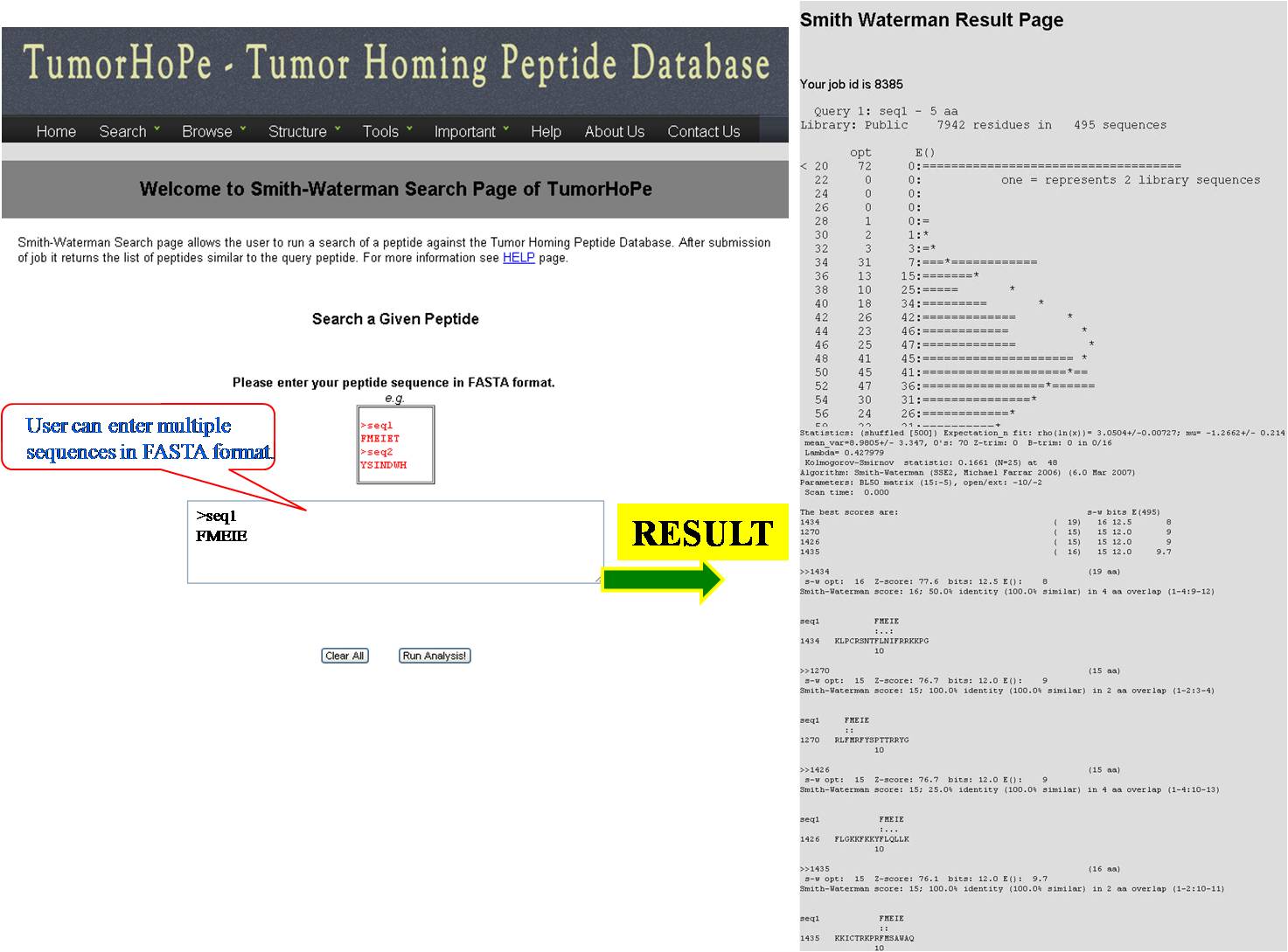
SMITH WATERMAN This tool can be used for determining similar regions between peptide sequences i.e. between query peptide and tumor homing peptides in the database, it compares the segments of possible lengths and optimizes the similarity measures giving exact result. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
IDENTICAL RESIDUES Here the query sequence will be mapped stepwise against all the peptides of Tumor Homing Peptide database and number of exact matched residues of each pair will give Identical Residues value as final output for that pair. more.. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
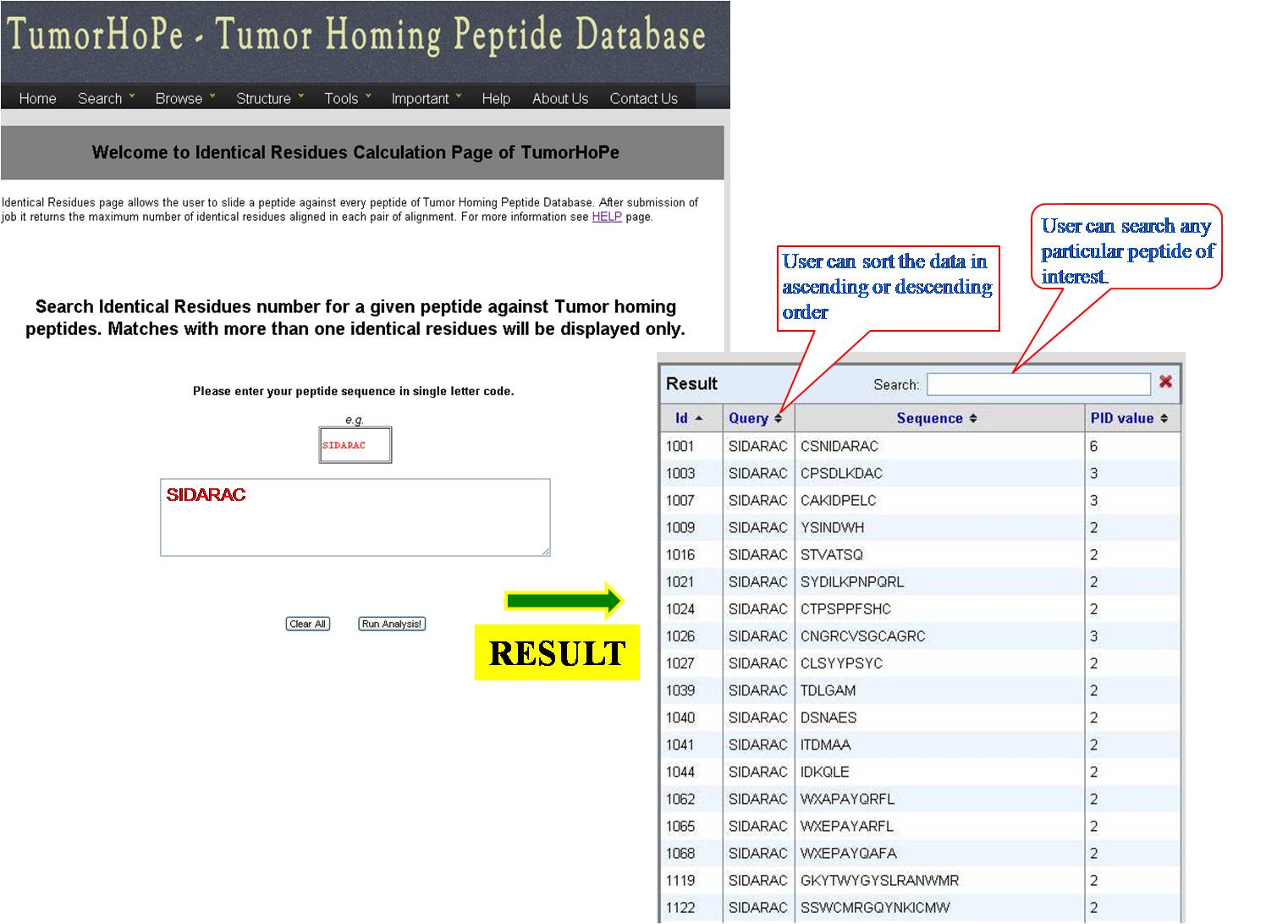 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
PEPTIDE MAPPING Peptide mapping is to find out if the query sequence or its motif is present in tumor homing peptide database. It is done in two ways-- SUBSEARCH: Maps the small peptide query against the tumor homing peptide database. SUPERSEARCH: Maps the tumor homing peptide database against large peptide/protein query. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
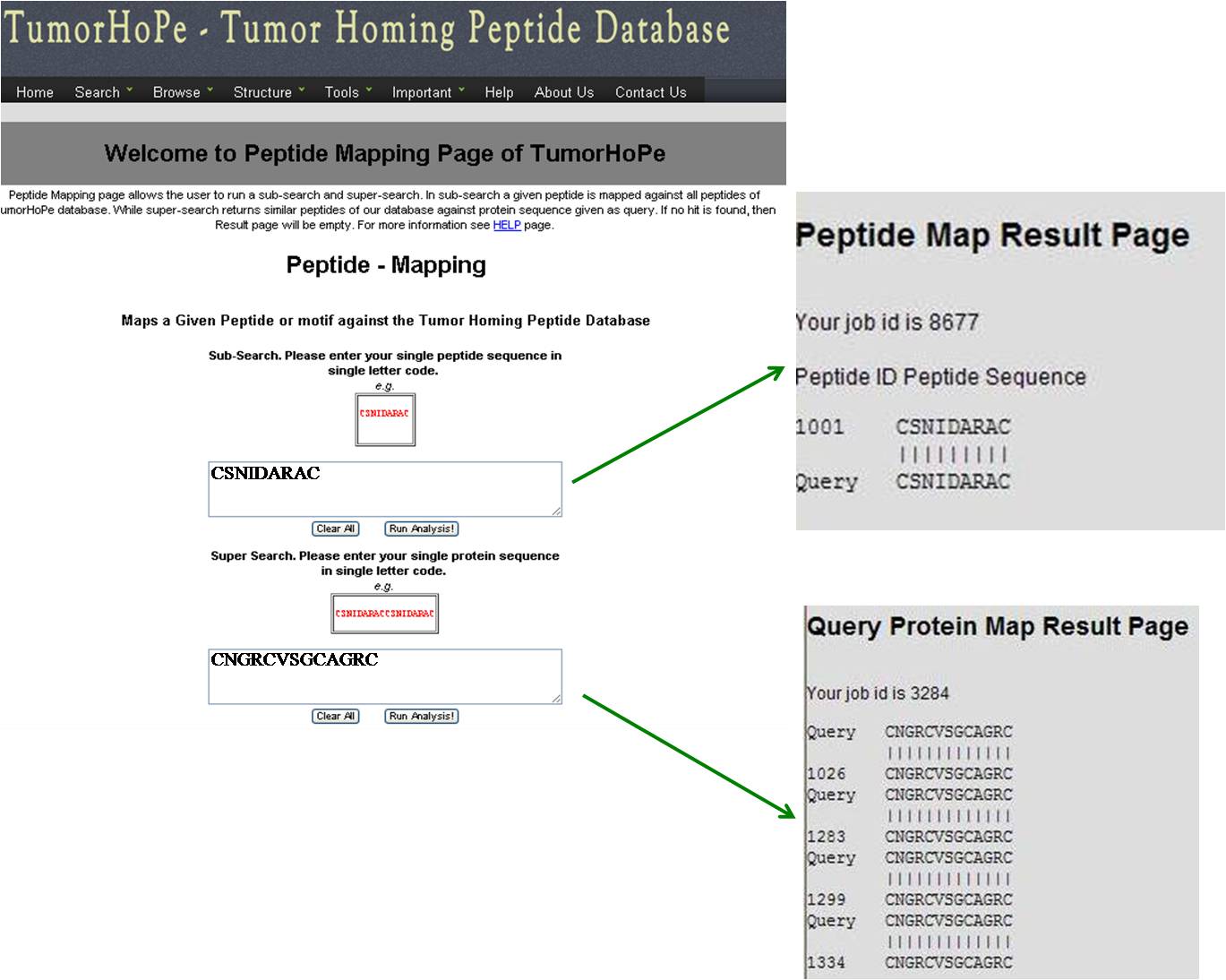 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
PHYSICAL PROPERTIES User gets physiochemical properties (hydrophobicity, hydrophilicity, Net charge, Iso electric point, Molecular weight) values for the query sequence. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
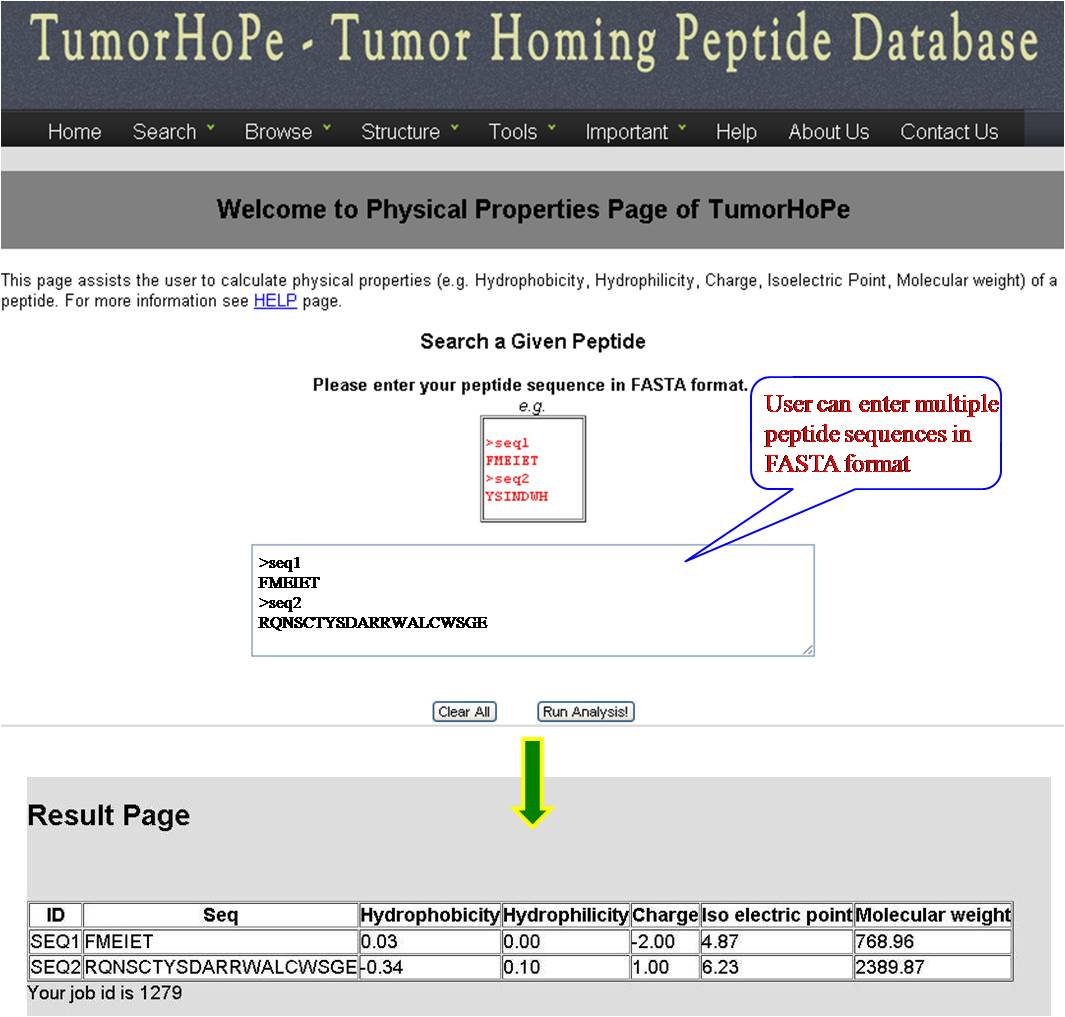 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DESCRIPTION OF TABLE AND FIELDS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
DESCRIPTION OF CELL LINES |
| Cell Line | ATCC No. | Organism | Disease | Source Oragnism | Morphology | Growth Properties |
|---|---|---|---|---|---|---|
| MDA-MB-231 | HTB-26 | Homo Sapiens | Adenocarcinoma | Mammary Gland | Epithelial | Adherent |
| MDA-MB-361 | HTB-27 | Homo Sapiens | Adenocarcinoma | Mammary Gland | Epithelial | Loosely Adherent |
| MDA-MB-435 | HTB-129 | Homo Sapiens | Ductal Carcinoma | Mammary Gland | Spindal | Adherent |
| MCF-7 | HTB-22 | Homo Sapiens | Adenocarcinoma | Mammary Gland | Epithelial | Adherent |
| SK-BR-3 | HTB-30 | Homo Sapiens | Adenocarcinoma | Mammary Gland | Epithelial | Adherent |
| HT-29 | HTB-38 | Homo Sapiens | Coloreactal Adenocarcinoma | Colon | Epithelial | Adherent |
| BT-474 | HTB-20 | Homo Sapiens | Ductal Carcinoma | Mammary Gland | Epithelial | Adherent, patchy |
| BT-483 | HTB-121 | Homo Sapiens | Ductal Carcinoma | Mammary Gland | Epithelial | Adherent |
| HeLa | CCL-2 | Homo Sapiens | Adenocarcinoma | Cervix | Epithelial | Adherent |
| Ca Ski | CRL-1550 | Homo Sapiens | Epidermoid Carcinoma | Cervix | Epithelial | Adherent |
| A549 | CCL-185 | Homo Sapiens | Carcinoma | Lung | Epithelial | Adherent |
| A375 | CRL_1619 | Homo Sapiens | Malignant Melanoma | Skin | Epithelial | Adherent |
| PC-3 | CRL_1435 | Homo Sapiens | Adenocarcinoma | Ovary | Epithelial | Adherent |
| MDA-PCa-2b | CRL-2422 | Homo Sapiens | Adenocarcinoma | Prostate | Epithelial | Adherent |
| LNCaP | CRL-1740 | Homo Sapiens | Carcinoma | Prostate | Epithelial | Adherent |
| OVCAR-3 | HTB-161 | Homo Sapiens | Adenocarcinoma | Ovary | Epithelial | Adherent |
| H226 | 5826 | Homo Sapiens | Squamous Cell Carcinoma | Lung | Epithelial | Adherent |
| H460 | HTB-177 | Homo Sapiens | Large cell lung cancer | Lung | Epithelial | Adherent |
| ZR-75-1 | CRL-1500 | Homo Sapiens | Ductal Carcinoma | Mammary Gland | Epithelial | Adherent |
| Hep G2 | HB-8065 | Homo Sapiens | Hepatocellular Carcinoma | Liver | Epithelial | Adherent |
| MIA PaCa-2 | CRL-1420 | Homo Sapiens | Carcinoma | Pancreas | Attached epithelial with floating rounded cells | Adherent |
Applications |
 |
 |
Frequently Asked Questions (FAQs) |
|
Q1. What is TumorHoPe? Ans . TumorHoPe is a literature based database of tumor homing peptides. Q2. Why tumor homing /targetting peptides? Ans. The biggest challenge of cancer chemotherapy is the lack of specificity and selectivity for the target. Peptide based therapy such as tumor homing peptides are highly specific for their target, these peptides target either tumor cells or microenvironment around them like bloodvessel and lymphvessel. Q3. How to search into TumorHoPe? Ans. User can search a peptide by name, peptide sequence, target tumor, cell lines and PMID. Q4. What other information one can get regarding peptide sequence? Ans. Data base is providing amino acid composition, frequency, other physiochemical properties like hydrophobicity, net charge along with this user can get secondary and tertiary structure information. Q5. Is this database useful if users have their own query sequence? Ans. Yes, user can use tools like BLAST, SMITH-WATERMAN, peptide mapping, they can also get information regarding amino acid composition, frequency and other physiochemical properties. Q6. Are all peptides used in the database experimentally validated? Ans. Most of the peptides are obtained by using phage display which give sets of clones for particular target cell, details are given about the clones having highest frequency of recovery. Q7. Is tumor homing peptide a reality? Ans. Yes, drugs are being conjugated with these peptides and are in different stages of clinical trials. e.g. tTF-NGR peptide was used in treatment of woman suffering from metastatic adenocarcinoma after 4 lines of chemotherapy. Q8. Are there any peptides with modification? Ans. Yes, there are few peptides with X amino acids, X represents norleucine, which is an isomer of leucine, the alpha amino acid 2-amino hexanoic acid. Also there are instances of sequences with F* where F* represents the 4-chlorophenylalanine. There are few amino acids with D-amino acids denoted by lower case letters. NOTE: If there is a biconjugate peptide with tumor killing peptide and homing peptide sequences, we have taken peptide sequence of homing peptide only in our database. |
