
- Home
- Predict
- Draw
- Analog Design
- Datasets
General ▾
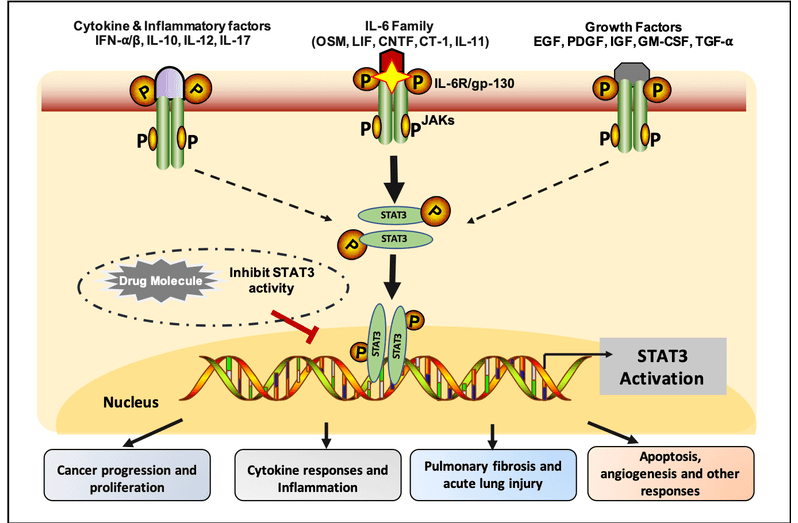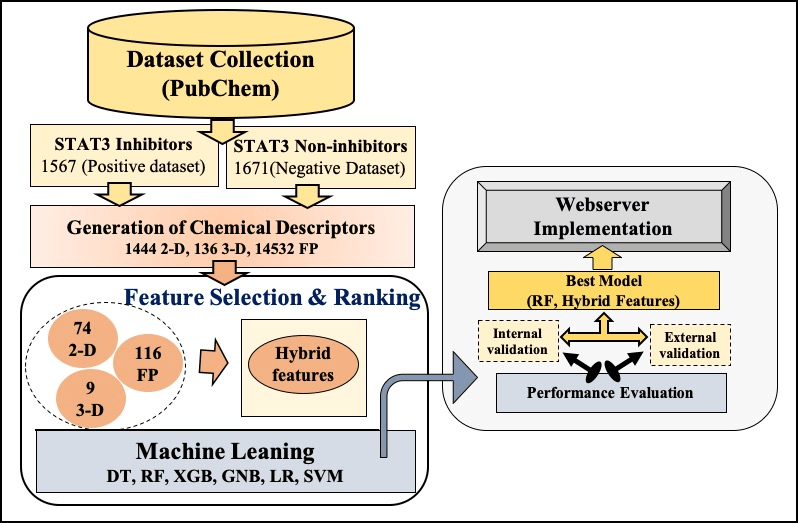
STAT3 is a pleiotropic transcription factor, which is activated in response to various cytokines such as IL6, IL10, and growth factors (FGF, EGF, and IGF). STAT3 mediated signaling pathway promotes the immunopathological responses, induction of inflammatory response and development of M2-like macrophages. STAT3 upregulation is associated with several pathological events, like cancer progression, proliferation, invasion, migration, angiogenesis, and multidrug resistance in various diseases. Additionally, STAT3 hyper-activation increases the cytokine storm production which play a major role in the pathogenesie ef COVID-19. Therefore, it is very important to design inhibitors which can supress the STAT3 activity, in order to cure a large number of diseases, associated with STAT3 activation. To facilitate researcher working in this era, we have developed a computational tool for the prediction and designing of STAT3 inhibitors/non-inhibitors.This server incorporate three modules to predict, draw, and design the chemical molecules which can inhibit STAT3 activation.