| |
siRNApred: Support vector machine based methods for predicting actual efficacy of both 21mer and 19mer siRNAs with high accuracy. SVM Methods were developed on two important datasets:
i) Main21 dataset consisting of 2182 siRNAs (21mer) derived from a homogeneous experimental conditions and its 19mer version, Main19 obtained by removing the two deoxyribonucleotide base-pair 3' overhang from above 21mer siRNA sequences
ii) Alternate19 dataset of 581 siRNAs (19mer) mined from seven different studies performed under heterogeneous experimental conditions.
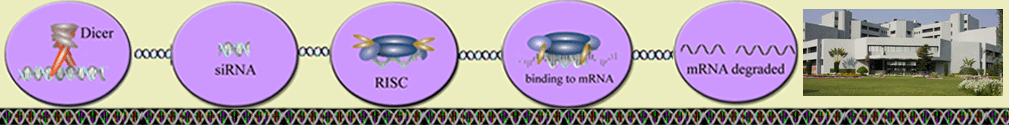
We have exploited nucleotide frequency feature i.e. occurrence of mono to penta nucleotide string in a siRNA sequence, to develop Hybrid-4 SVM method. Binary patterns i.e. presence or absence of A, G, C, T/U nucleotides at each position in a siRNA sequence were used to develop Binary pattern SVM method and finally combination of both approaches to develop Hybrid-7 SVM method.
|
|